Linking remote contigs to isolate records¶
If remote contigs have been enabled, isolates can be linked to contigs stored in an external BIGSdb database, rather than directly uploaded. These well then be loaded when needed, for example during scanning or data export. This will be marginally slower than hosting contigs within the same database, but minimises duplication of sequence data and associated storage. Contigs need to be accessible via the BIGSdb RESTful API.
Click the sequences link icon on the curator’s main page.
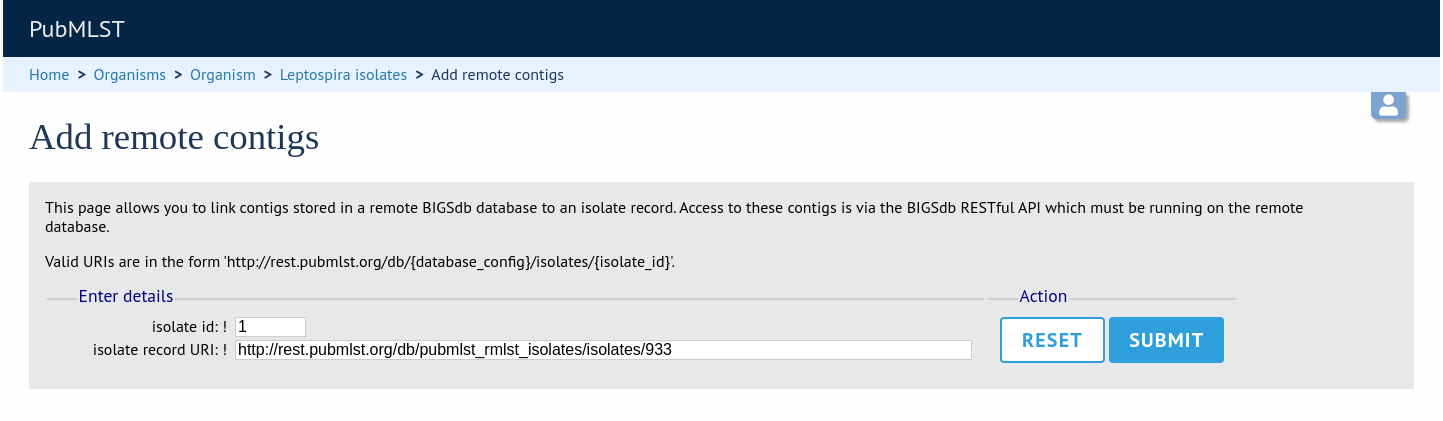
Either select the isolate id from the dropdown list, or enter it manually (list is disabled if there are >1000 records in the database). Enter the URI for the RESTful API of the parent isolate record, e.g. http://rest.pubmlst.org/db/pubmlst_rmlst_isolates/isolates/933. This URI can require authentication if credentials have been set up.
Press submit.
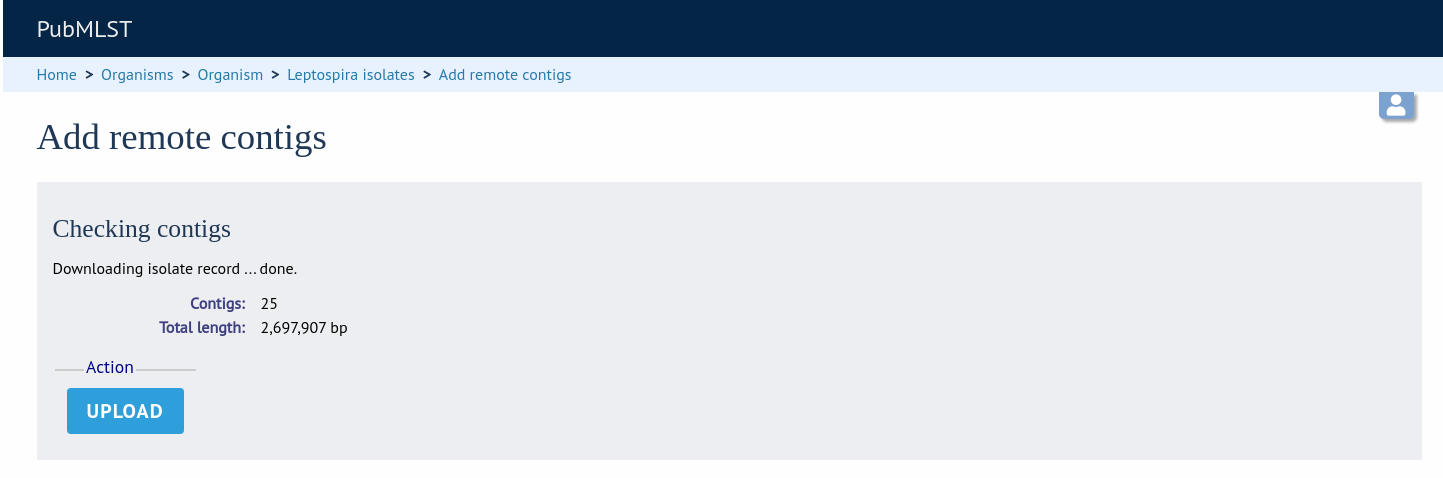
Summary information about the number of contigs and their total length will be downloaded from the remote isolate record. You will then be prompted to upload this information to the database, by clicking the ‘Upload’ button.
The contigs will be downloaded in bulk in order to determine their lengths. This information is stored within the local database as it is required for various outputs. Full metadata is not stored at this stage.
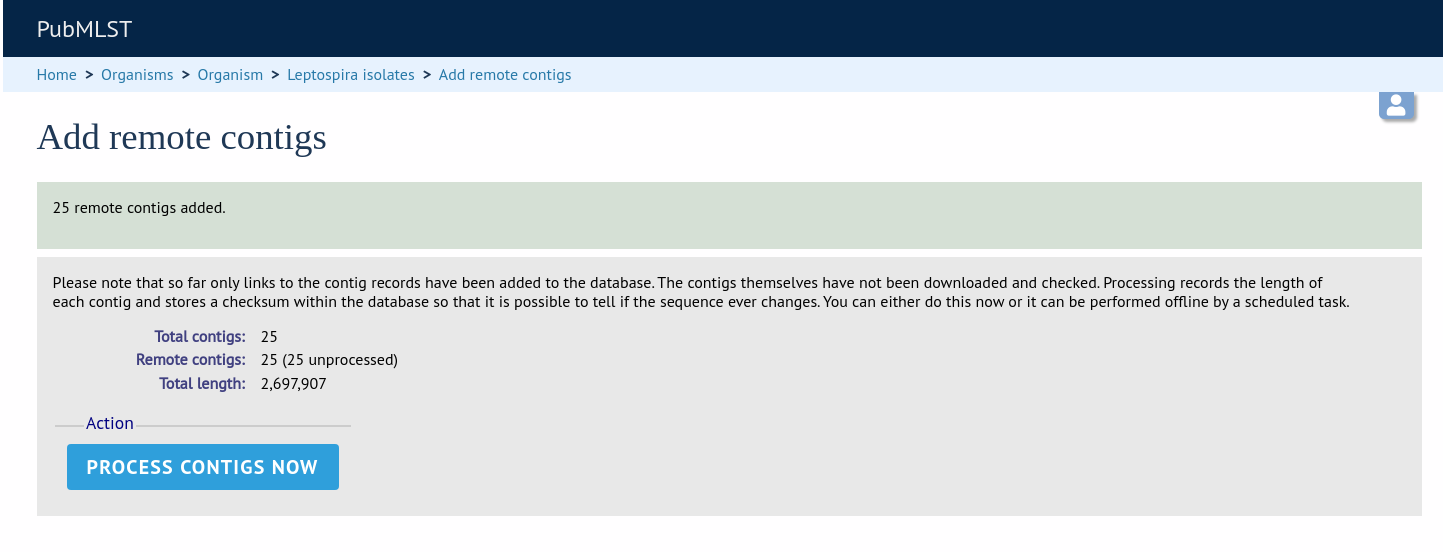
This is all that is required for the contigs to be used as normal. In order to get the full metadata about the contigs (sequencing platform used, sender and datestamp information), you can choose to process the contigs by clicking the ‘Process contigs now’ button. This will download each contig in turn, and store its provenance metadata locally.
Alternatively, this step can be performed offline automatically.