Contig export
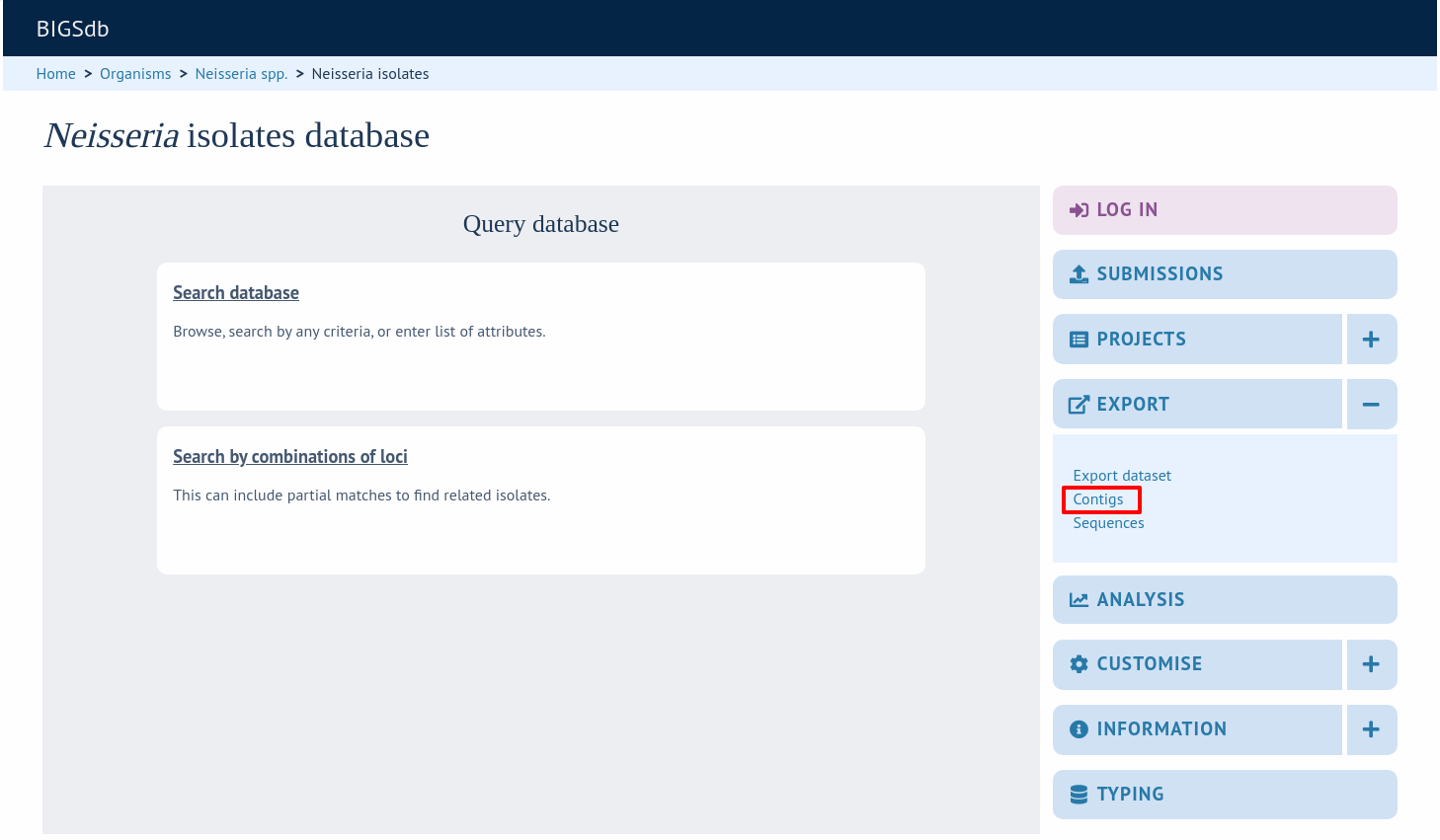
The contig export plugin can be accessed by expanding the ‘Export’ section and clicking the ‘Contigs’ link in the contents page of isolate databases.
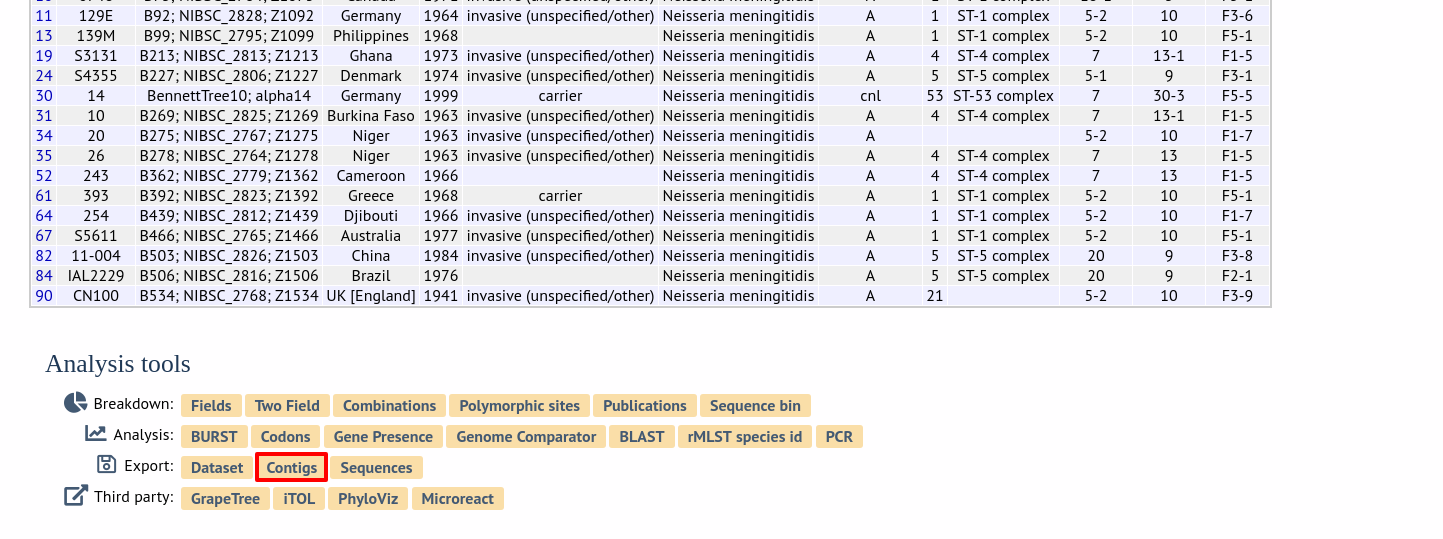
Alternatively, it can be accessed following a query by clicking the ‘Contigs’ button in the Export section at the bottom of the results table.
Select the isolates for which you wish to export contig data for. In databases with a large number of isolates you will need to enter the id numbers rather than select from a list. If the export function was accessed following a query, isolates returned in the query will be pre-selected.
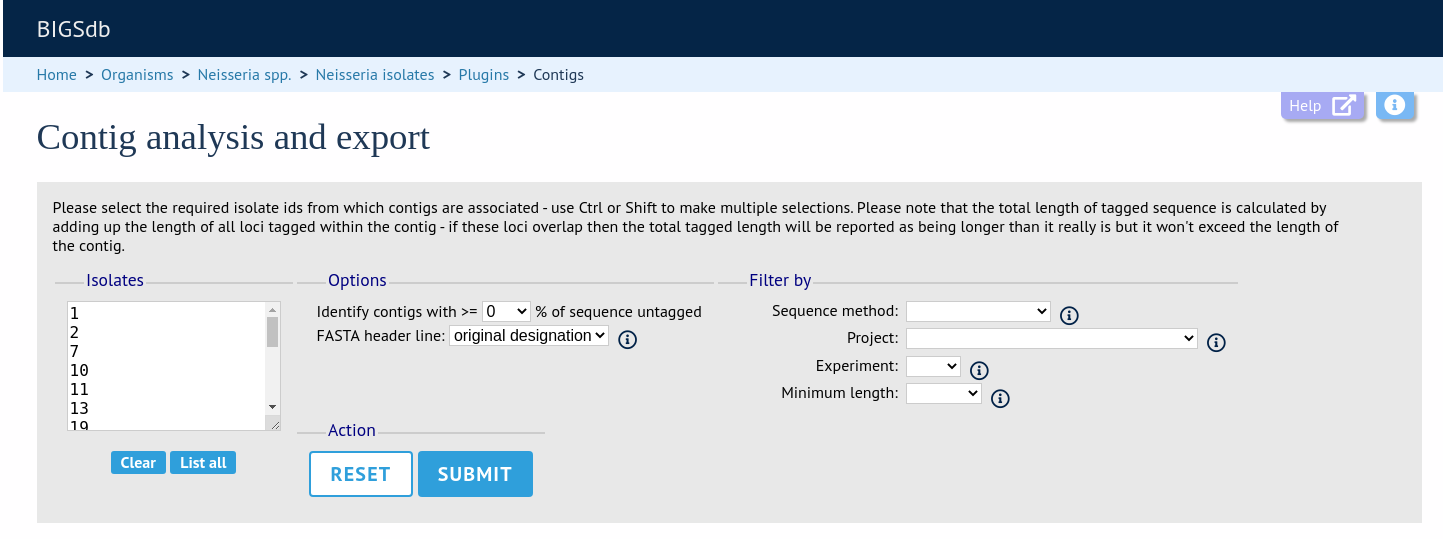
At its simplest, press submit.
A table will be produced with download links. Clicking these will produce the contigs in FASTA format.
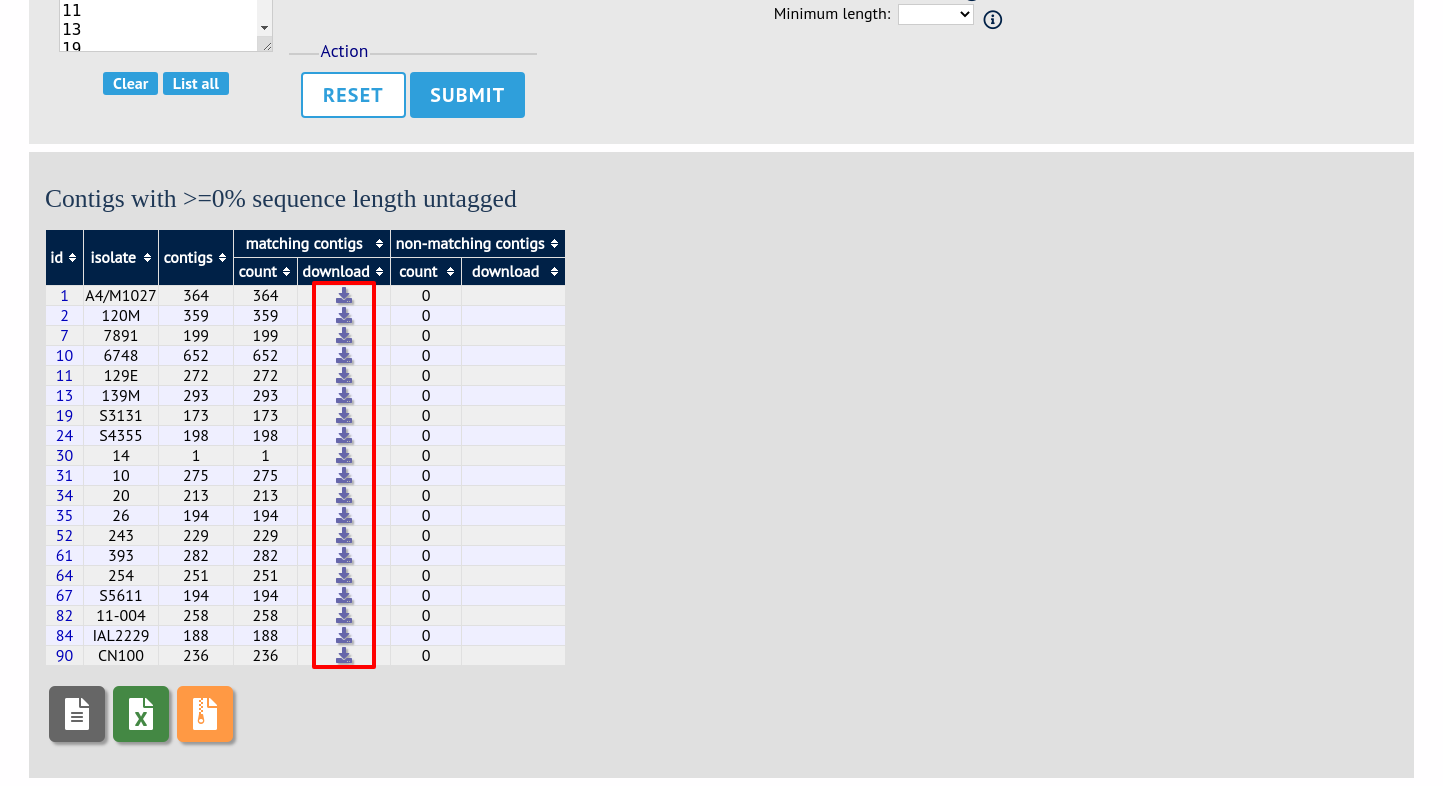
You can also download all the data in a tar file by clicking the ‘Batch download’ link.
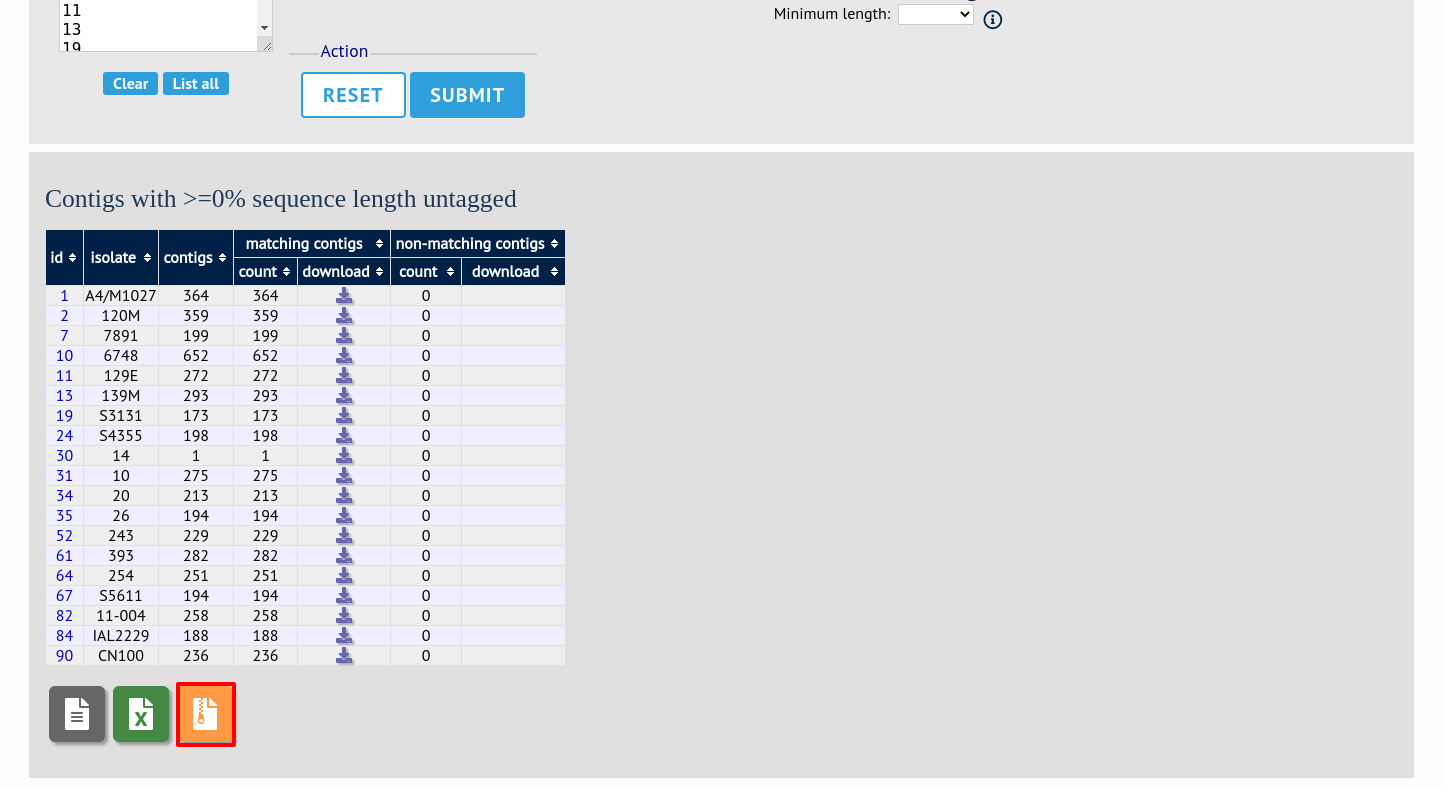
Filtering by tagged status of contigs
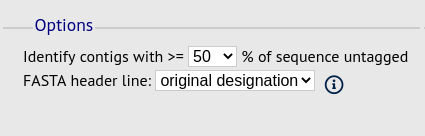
You can also export contigs based on the percentage of the sequence that has been tagged. This is useful to find sequences to target for gene discovery.
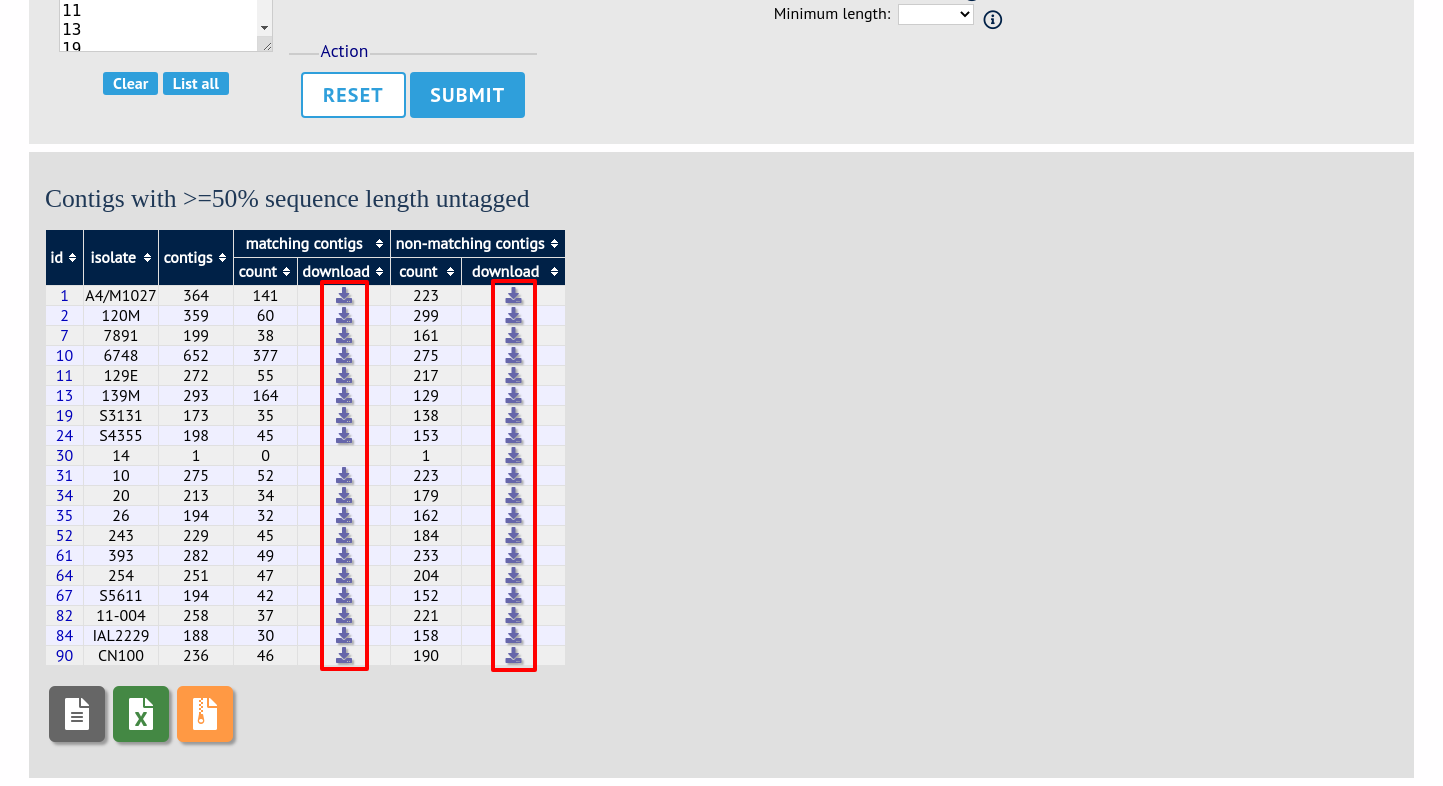
In order to export contigs where at least half the sequence has been tagged (and also the remaining contigs in a separate file), select ‘50’ in the dropdown box for %untagged.
The resulting table has two download links for each isolate, one for contigs matching the condition, and one for contigs that don’t match.