Querying sequences to determine allele identity
Sequence queries are performed in the sequence definition database. Click ‘Single sequence’ query from the contents page.
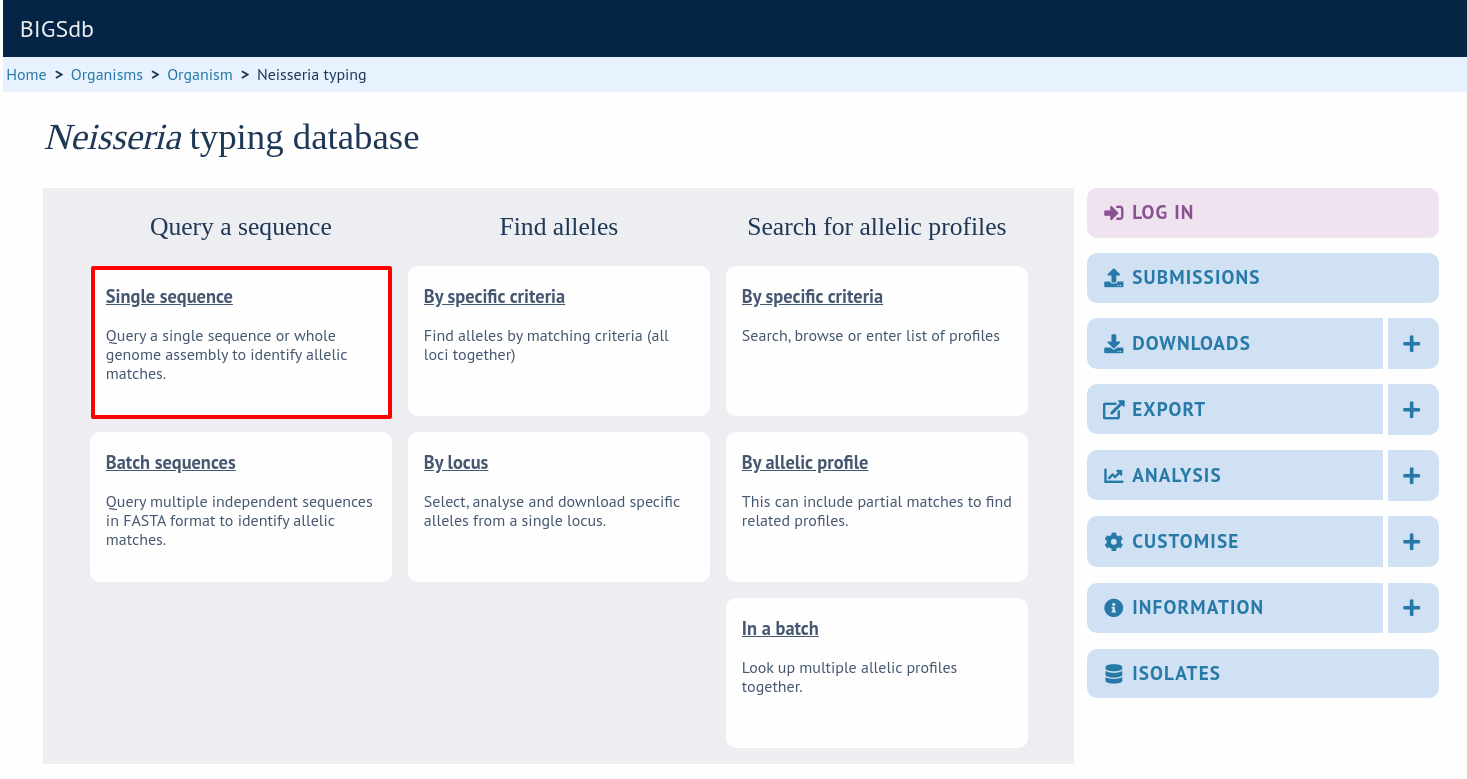
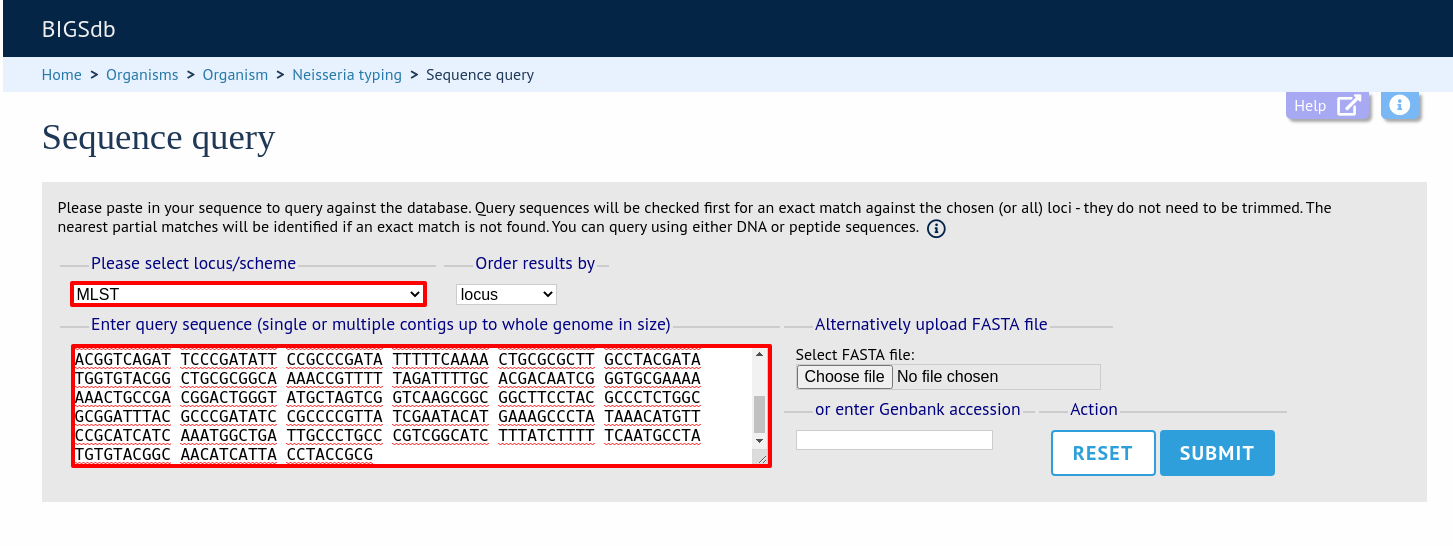
Paste your sequence in to the box - there is no need to trim. Often, you can leave the locus setting on ‘All loci’ - the software should identify the correct locus based on your sequence. Sometimes, especially in databases with a large number of defined loci, it may be quicker, however, to select the specific locus or scheme (e.g. MLST) that a locus belongs to.
Note
If the locus you are querying is a shorter version of another, e.g. an MLST fragment of a gene where the full length gene is also defined, you will need to select the specific locus or the scheme from the dropdown box. Leaving the selection on ‘All loci’ will return a match to the longer sequence in preference to the shorter one.
Click ‘Submit’.
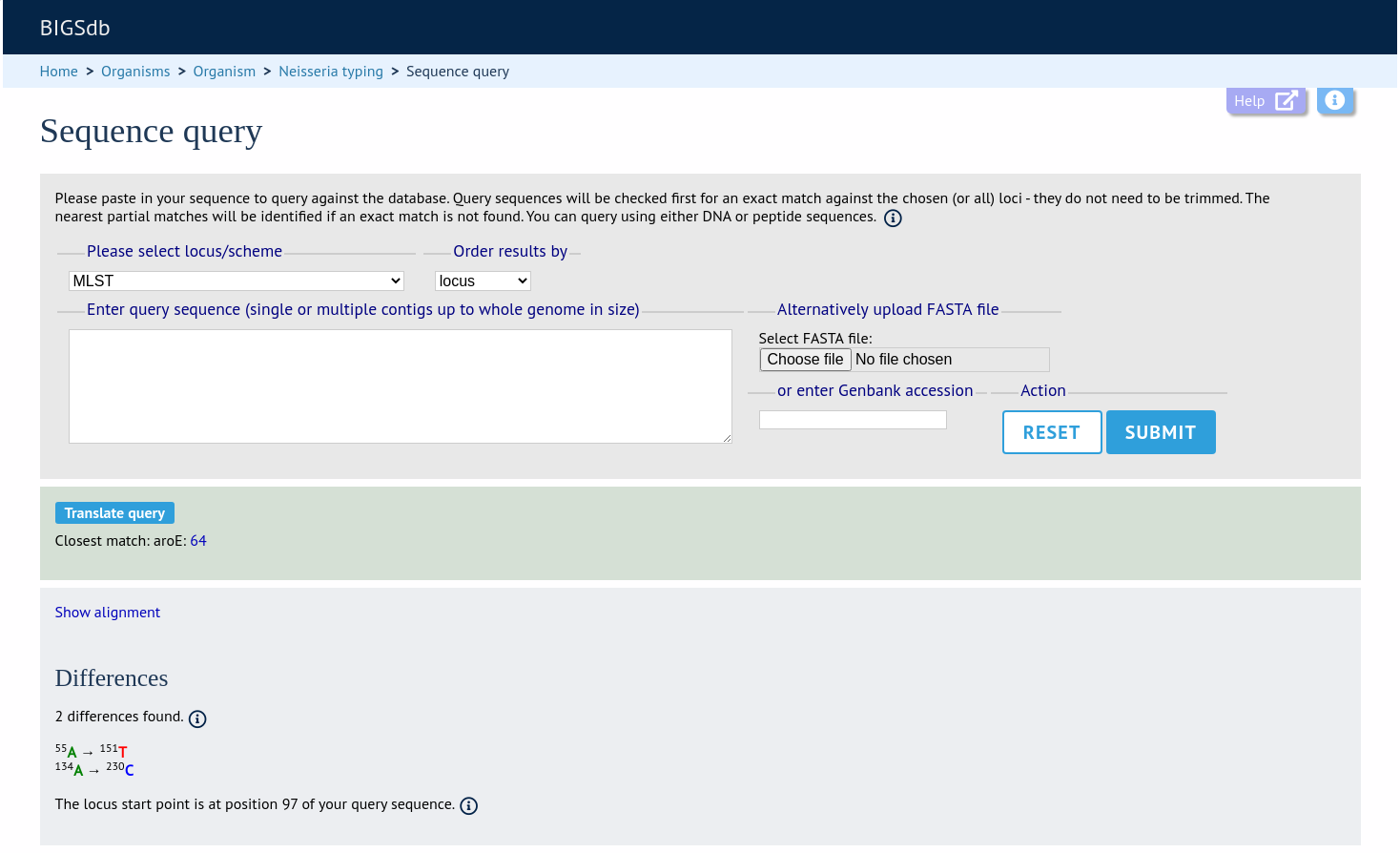
If an exact match is found, this will be indicated along with the start position of the locus within your sequence.
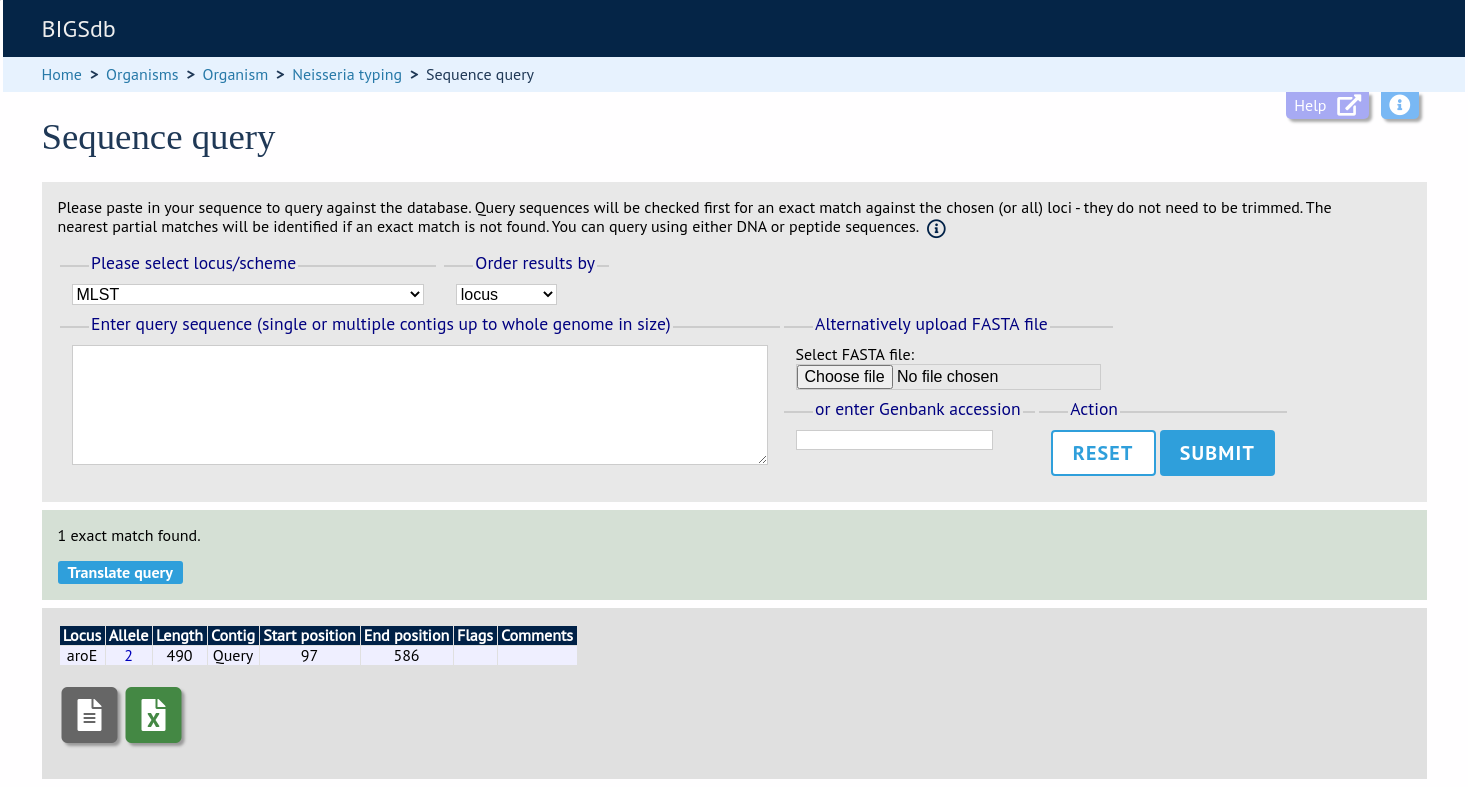
If only a partial match is found, the most similar allele is identified along with any nucleotide differences. The varying nucleotide positions are numbered both relative to the pasted in sequence and to the reference sequence. The start position of the locus within your sequence is also indicated.
As an alternative to pasting a sequence in to the box, you can also choose to upload sequences in FASTA format by clicking the file browse button.
Querying whole genome data
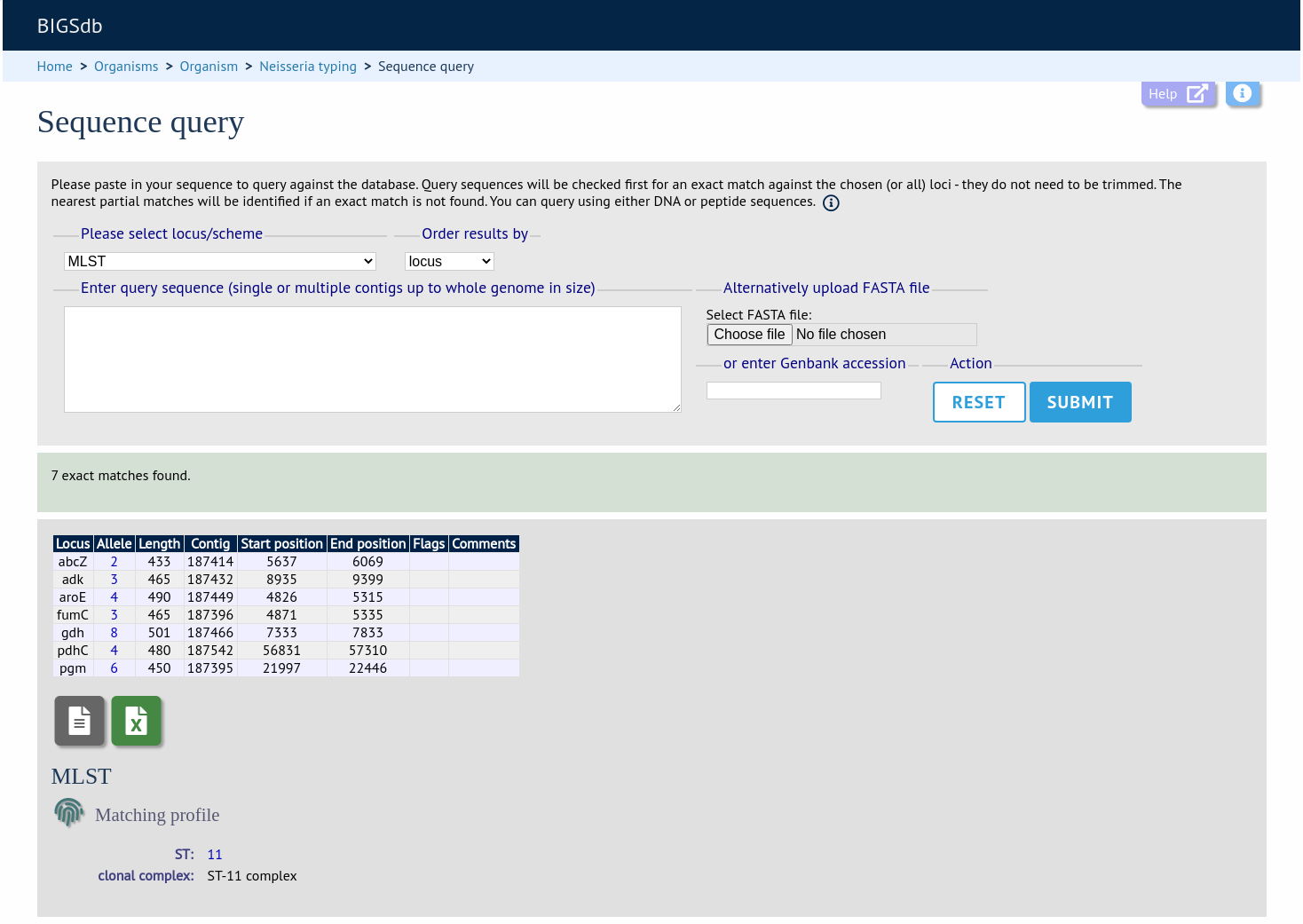
The sequence query is not limited to single genes. You can also paste or upload whole genomes - these can be in multiple contigs. If you select a specific scheme from the dropdown box, all loci belonging to that scheme will be checked (although only exact matches are reported for a locus if one of the other loci has an exact match). If all loci are matched, scheme fields will also be returned if these are defined. This, for example, enables you to identify the MLST sequence type of a genome in one step.
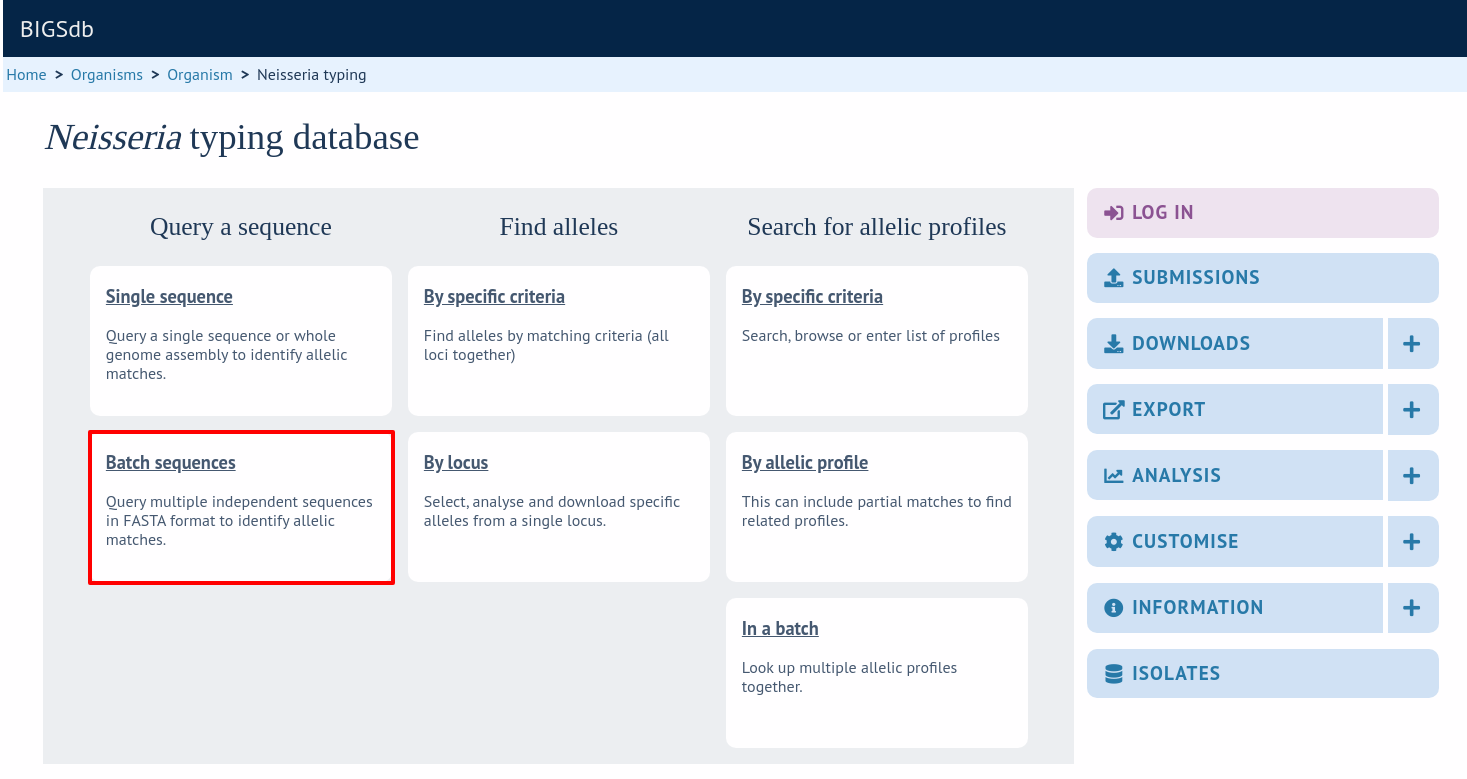
Querying multiple sequences to identify allele identities
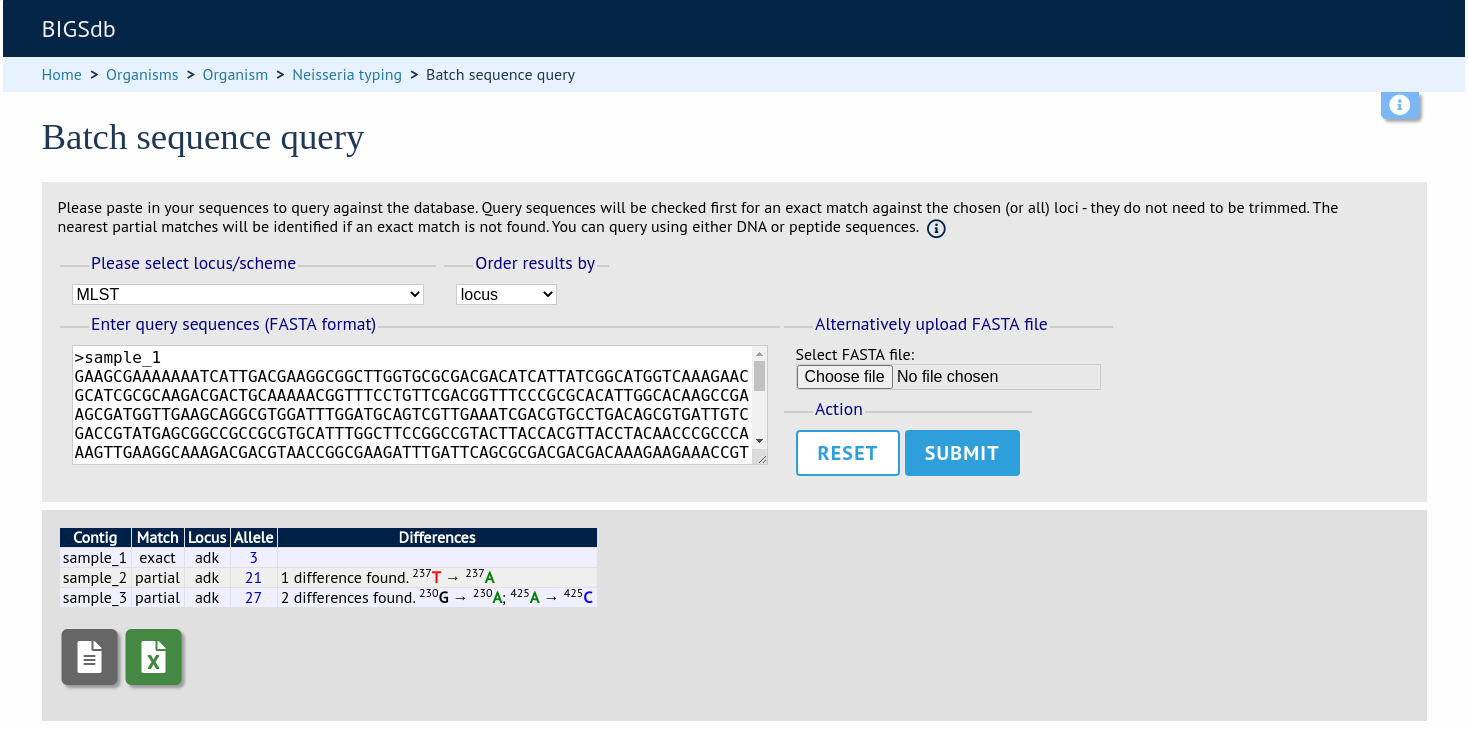
You can also query mutiple sequences together. These should be in FASTA format. Click ‘Batch sequences’ from the contents page.
Paste your sequences (FASTA format) in to the box. Select a specific locus, scheme or ‘All loci’.
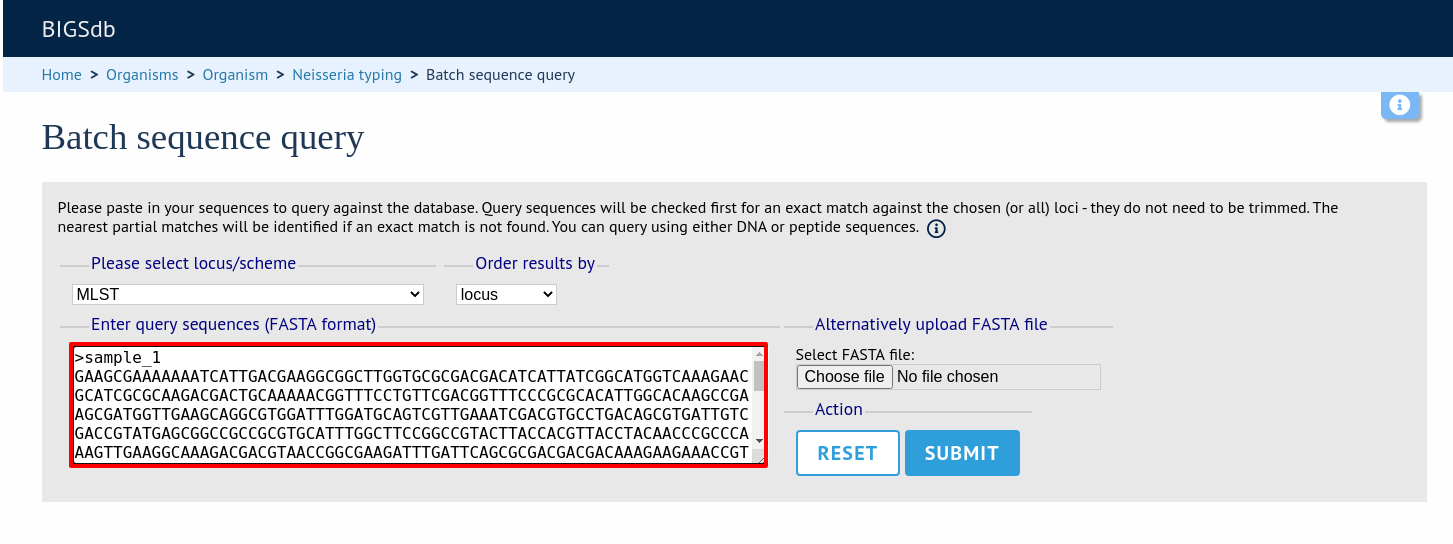
The best match will be displayed for each sequence in your file. If this isn’t an exact match, the differences will be listed.