In silico PCR
This is a tool that can be used to simulate PCR reactions run against genomes stored in the database. This is useful for designing and testing primers. The plugin uses the exonerate ipcress program to perform its simulation.
The tool can be accessed by selecting the ‘Analysis’ section on the main contents page.
Jump to the ‘Analysis’ category, follow the link to BLAST, then click ‘Launch PCR’.

Alternatively, it can be accessed following a query by clicking the ‘PCR’ button at the bottom of the results table. Isolates returned from the query will be automatically selected within the analysis interface.
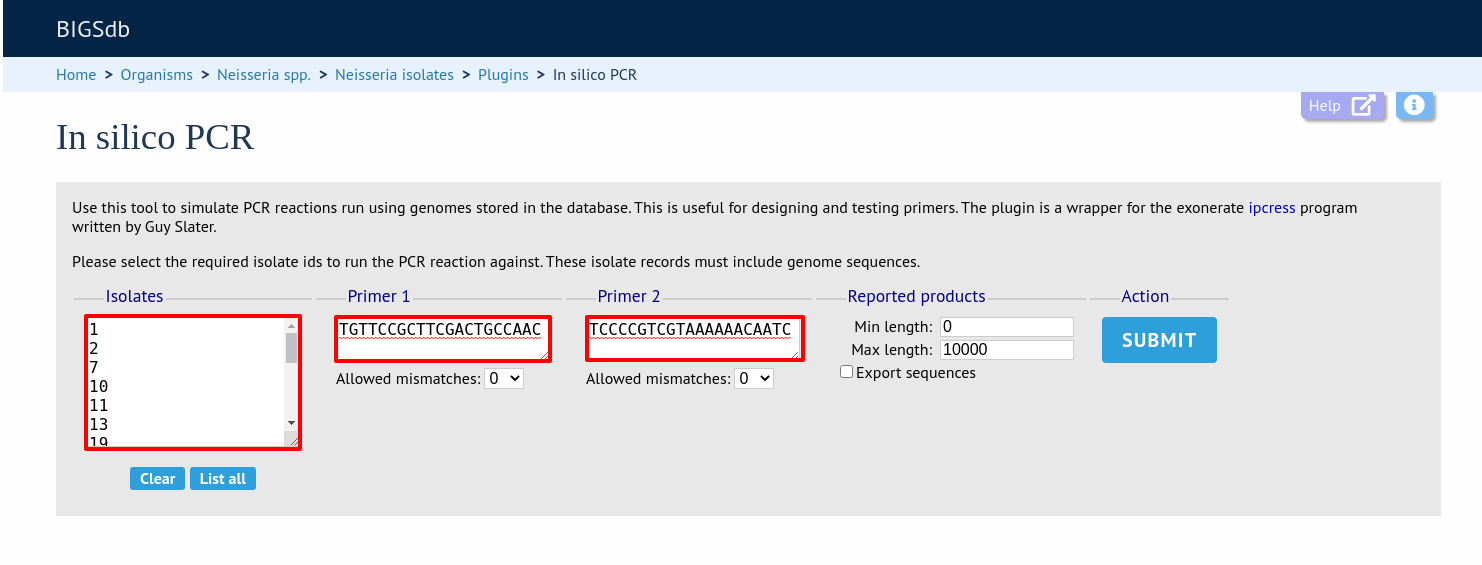
Select the isolates to include. These will be pre-populated if you arrive here following a search.
Enter your forward and reverse primer sequences in the appropriate boxes. These may contain wobble bases if necessary. You can also specify how many mismatches are allowed for each primer. Finally, you can restrict the reported length to only those products that fall between a minimum and maximum length range.
Click ‘Submit’. The job will be sent to the job queue. The output will be a table of predicted products, showing the number of products and their positions within a contig. A summary of this table is also availabe to download in tab- delimited text of Excel formats.
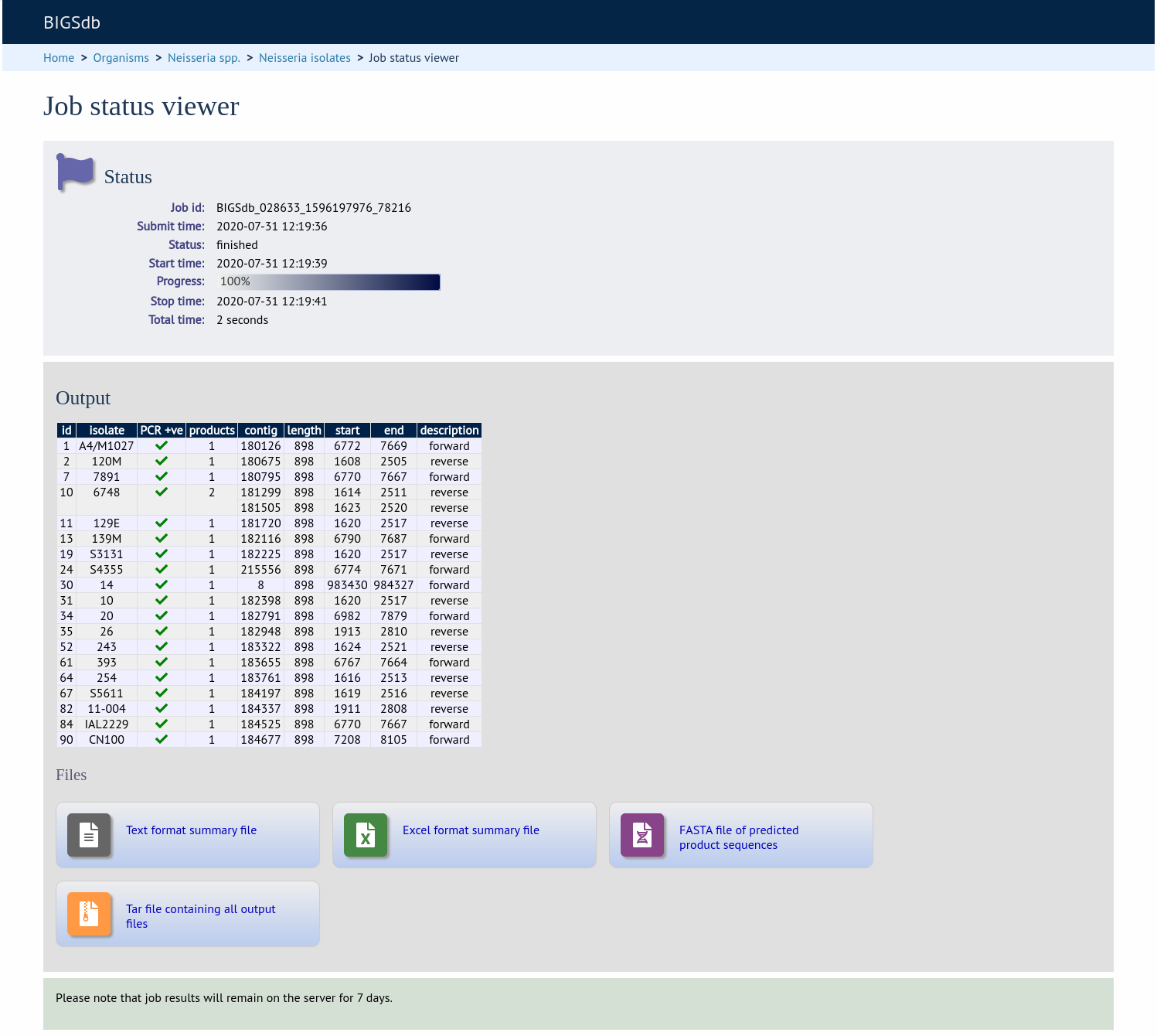
It is also possible to export the predicted product sequence. You can do this by selecting the ‘Export sequences’ checkbox on the options form.
Note
The exported sequences will include the primer regions. It is important to note that, unlike a real PCR reaction, these sequences represent the sequence within this region in the genome. In a real PCR reaction, the primers are themseleves incorporated in to the product, so even if there was a mismatch in the primer region, the product sequence would include the primer sequence.