Kleborate
Kleborate is a tool that can be used to screen assemblies of Klebsiella pneumoniae and the Klebsiella pneumoniae species complex (KpSC) for:
MLST sequence type
species (e.g. K. pneumoniae, K. quasipneumoniae, K. variicola, etc.)
ICE Kp associated virulence loci: yersiniabactin (ybt), colibactin (clb), salmochelin (iro), hypermucoidy (rmpA)
virulence plasmid associated loci: salmochelin (iro), aerobactin (iuc), hypermucoidy (rmpA, rmpA2)
antimicrobial resistance determinants: acquired genes, SNPs, gene truncations and intrinsic β-lactamases
K (capsule) and O antigen (LPS) serotype prediction, via wzi alleles and Kaptive.
Kleborate and Kaptive are described in Lam et al. 2021 Nat Commun 12: 4188 (PubMed: 34234121) and Lam et al. 2022 Microb Genom 8: 000800 (PubMed: 35311639) respectively.
Kleborate will usually only be available on databases hosting suitable data i.e. Klebsiella isolates.
The function can be accessed by selecting the ‘Analysis’ section on the main contents page.
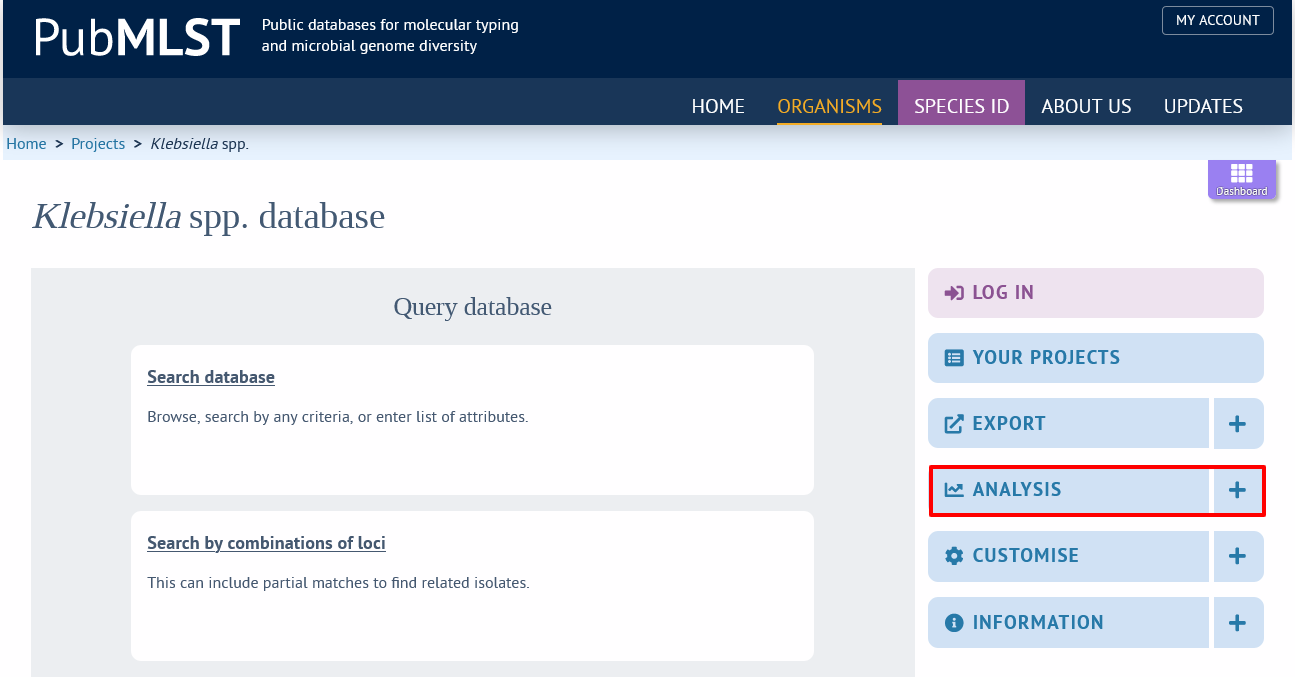
Jump to the ‘Third party’ category, follow the link to Kleborate, then click ‘Launch Kleborate’.

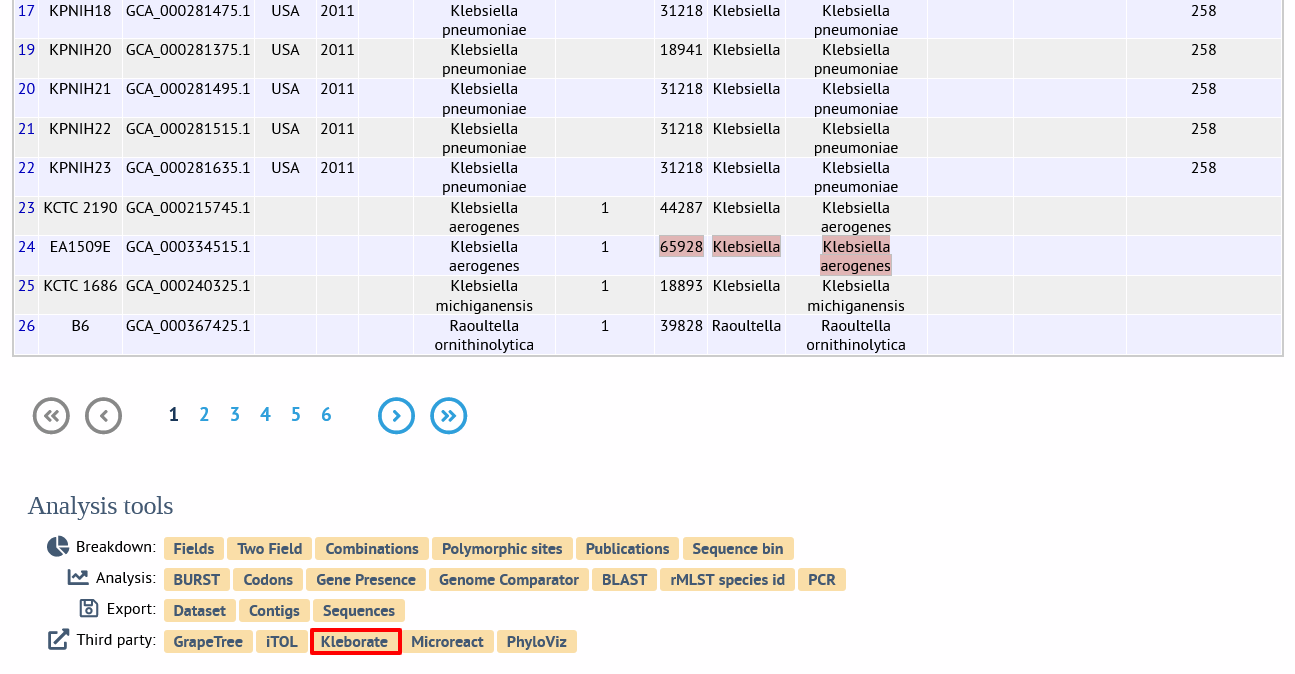
Alternatively, it can be accessed following a query by clicking the ‘Kleborate’ button in the ‘Third party’ list at the bottom of the results table. Please note that the list of functions here may vary depending on the setup of the database.
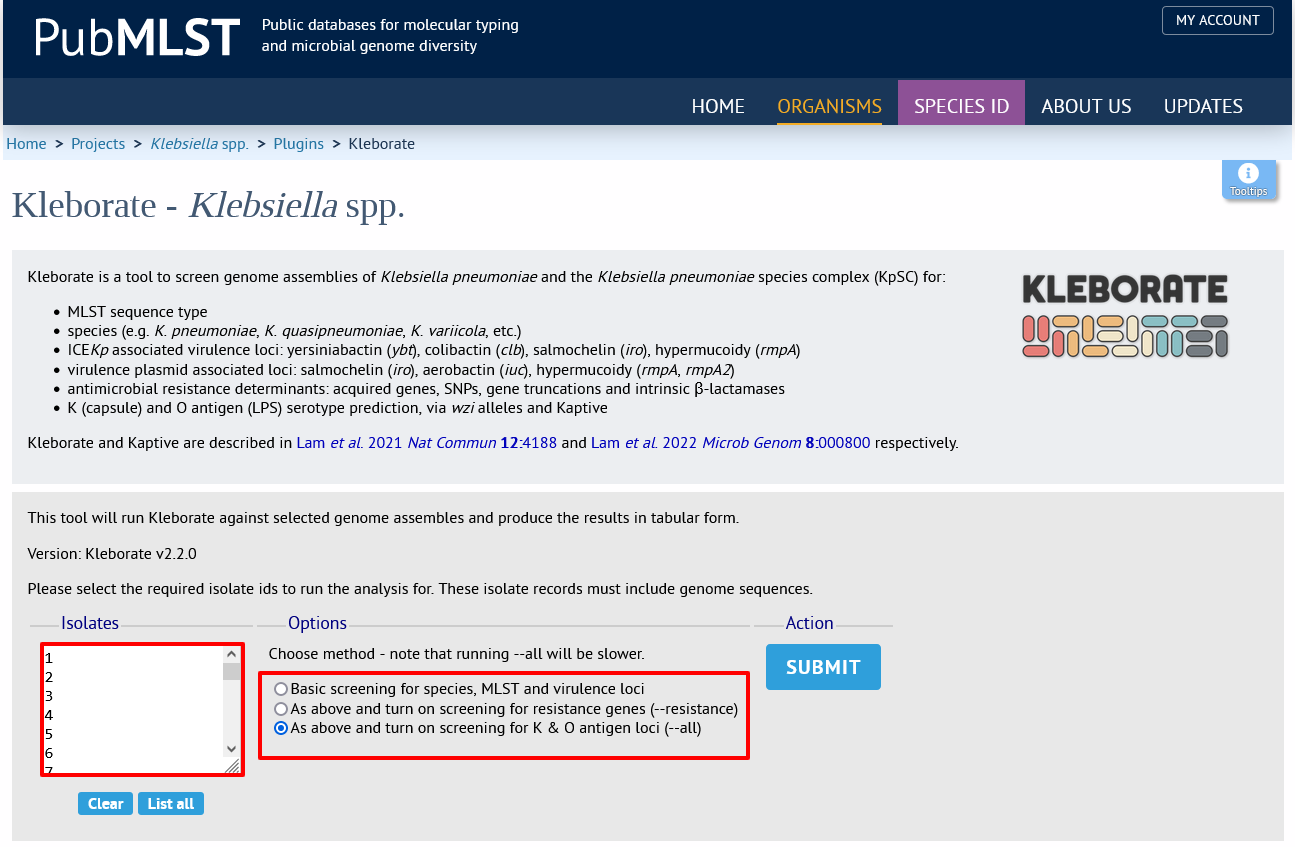
Select the isolate records to analyse - these will be pre-selected if you accessed the plugin following a query. You can then choose the analysis to run:
Basic screening for species, MLST and virulence loci
As above and turn on screening for resistance genes (–resistance)
As above and turn on screening for K & O antigen loci (–all)
Click submit.
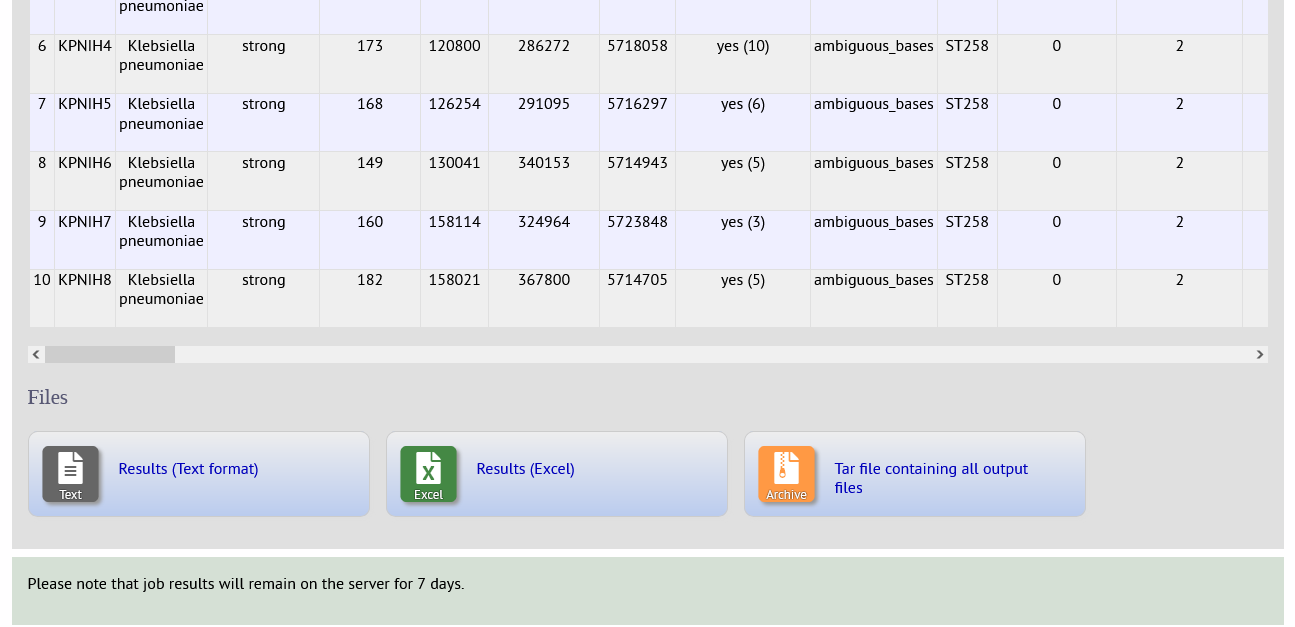
The analysis will be submitted to the job queue.
When the job has completed you will see a table of results that is also available for download in tab-delimited text and Excel formats.