Interactive Tree of Life (iTOL)
The ITOL plugin allows you to generate and visualise phylogenetic trees calculated from concatenated sequence alignments of selected loci (or the loci belonging to a particular scheme). Currently, only Neighbour-joining trees are supported. Datasets can include metadata which allows nodes in the resultant tree to be coloured.
ITOL can be accessed by selecting the ‘Analysis’ section on the main contents page.
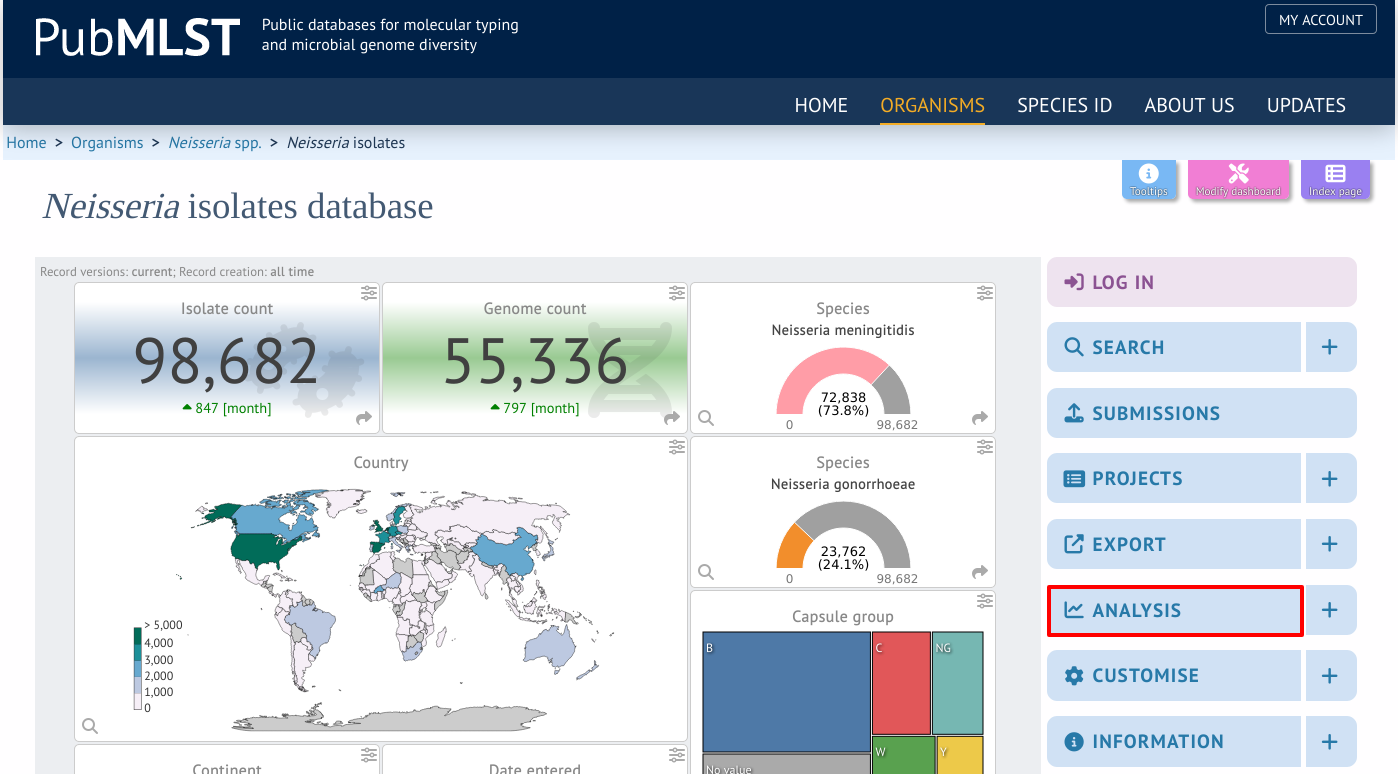
Jump to the ‘Third party’ category, follow the link to iTOL, then click ‘Launch iTOL’.
ITOL can be accessed from the contents page by clicking the ‘iTOL’ link.
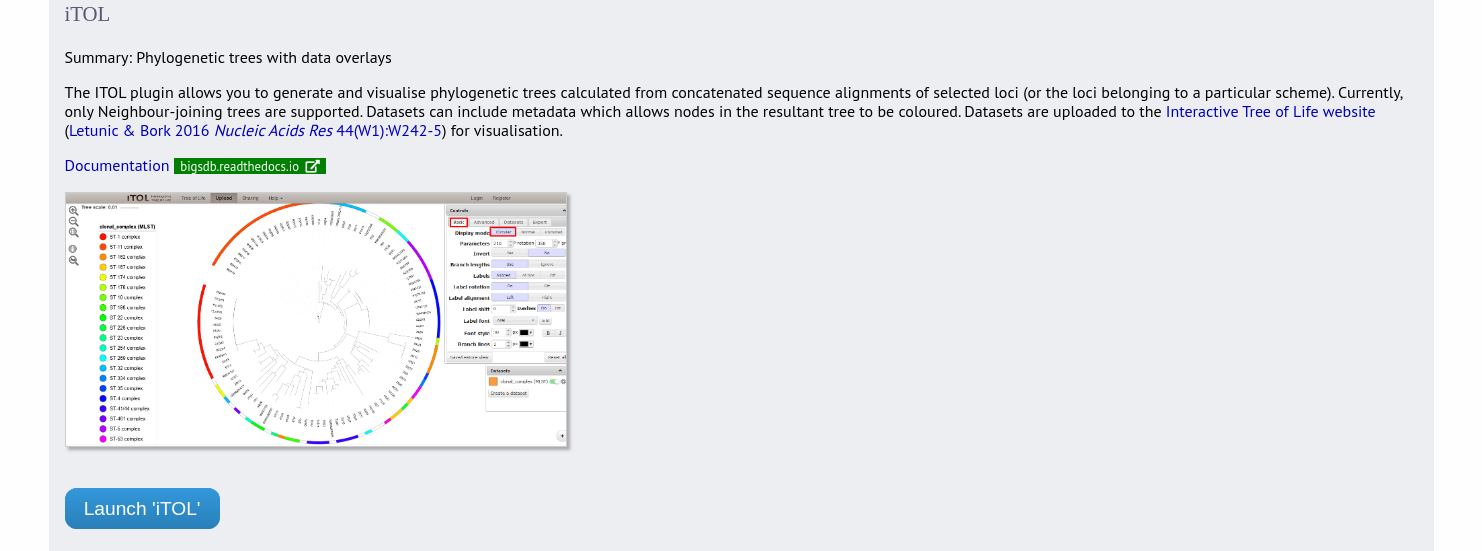
Alternatively, it can be accessed following a query by clicking the ‘iTOL’ button at the bottom of the results table. Isolates returned from the query will be automatically selected within the iTOL interface.
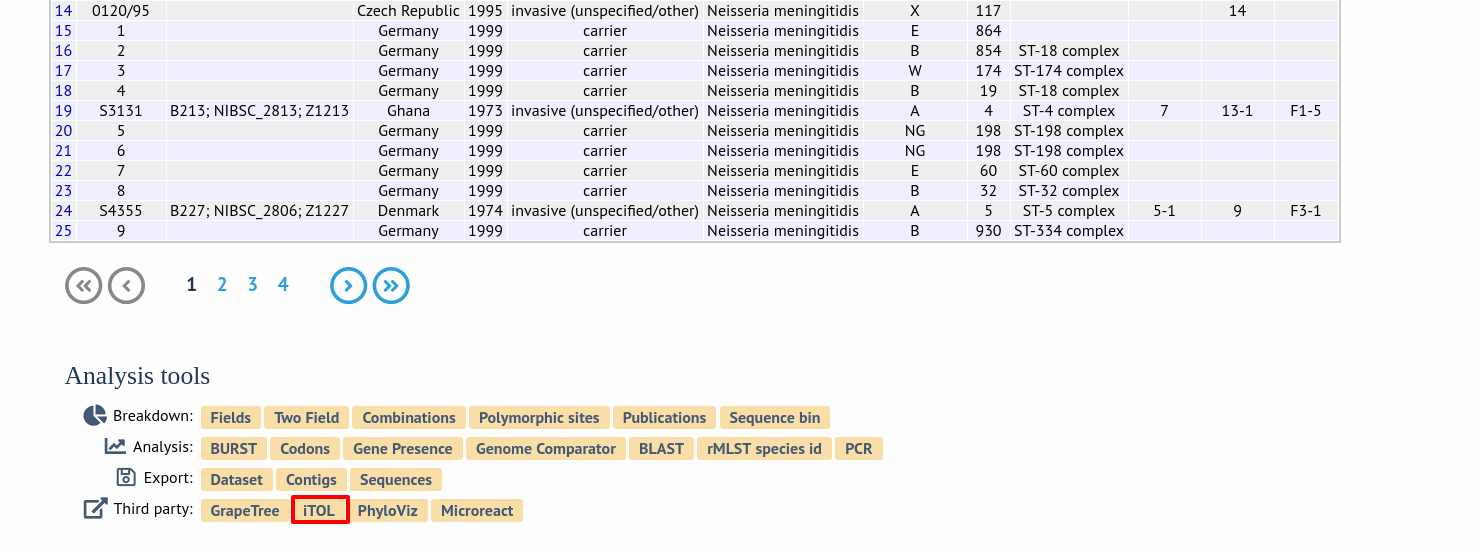
Select the isolates to include. The tree can be generated from concatenated sequences of any selection of loci, or more conveniently, you can select a scheme in the scheme selector, or in the list of recommended schemes if these have been set up, to include all loci belonging to that scheme.
Additional fields can be selected to be included as metadata for use in colouring nodes - select any fields you wish to include in the ‘iTOL datasets’ list. Multiple selections can be made by holding down Shift or Ctrl while selecting. You can also choose how nodes are labeled by metadata - either by colouring the labels or using coloured strips.
Click ‘Submit’ to start the analysis.

The job will be sent to the job queue. When it has finished, the generated tree and associated metadata will be uploaded to the Interactive Tree of Life website (https://itol.embl.de/). Click the button marked ‘Launch iTOL’.
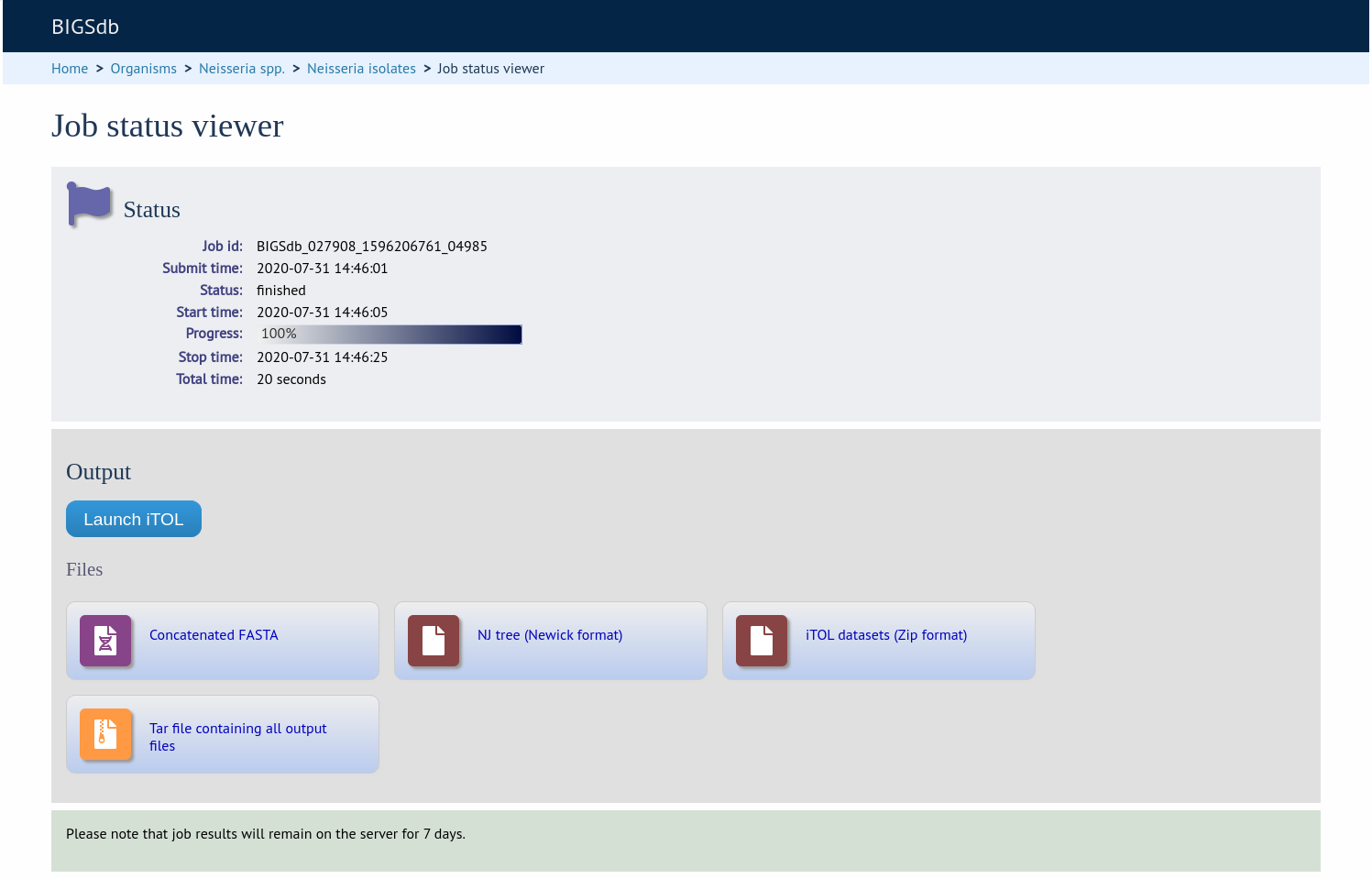
Your browser will open the iTOL website with your tree.
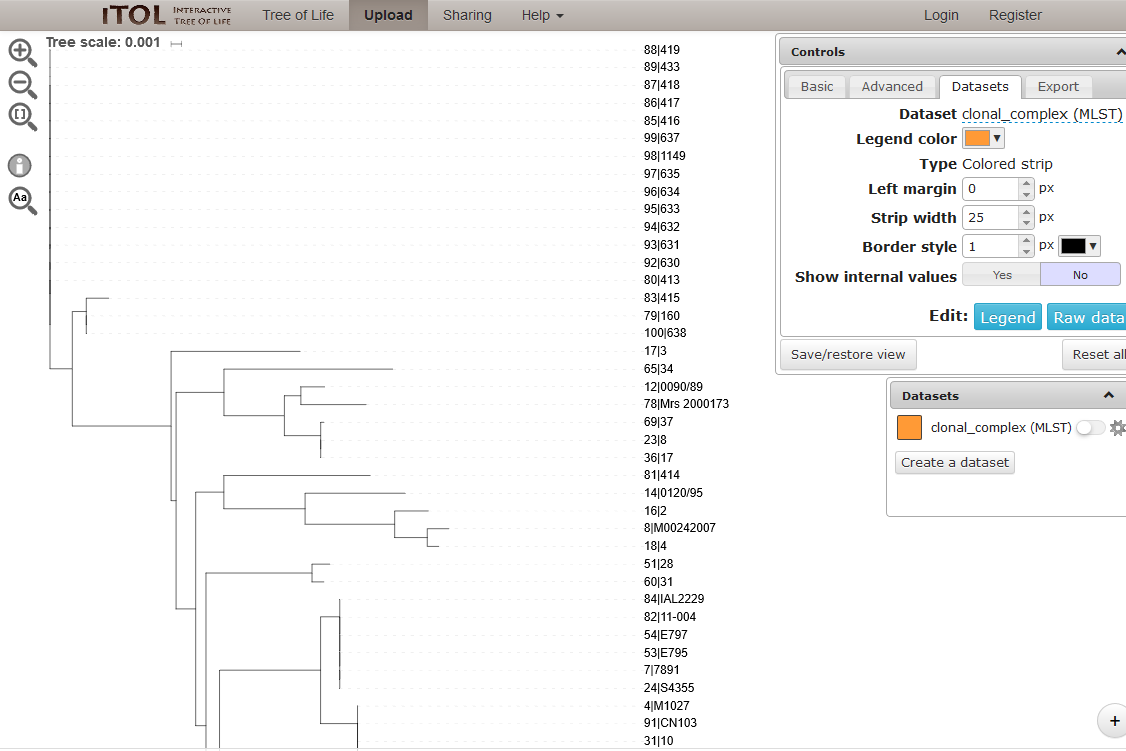
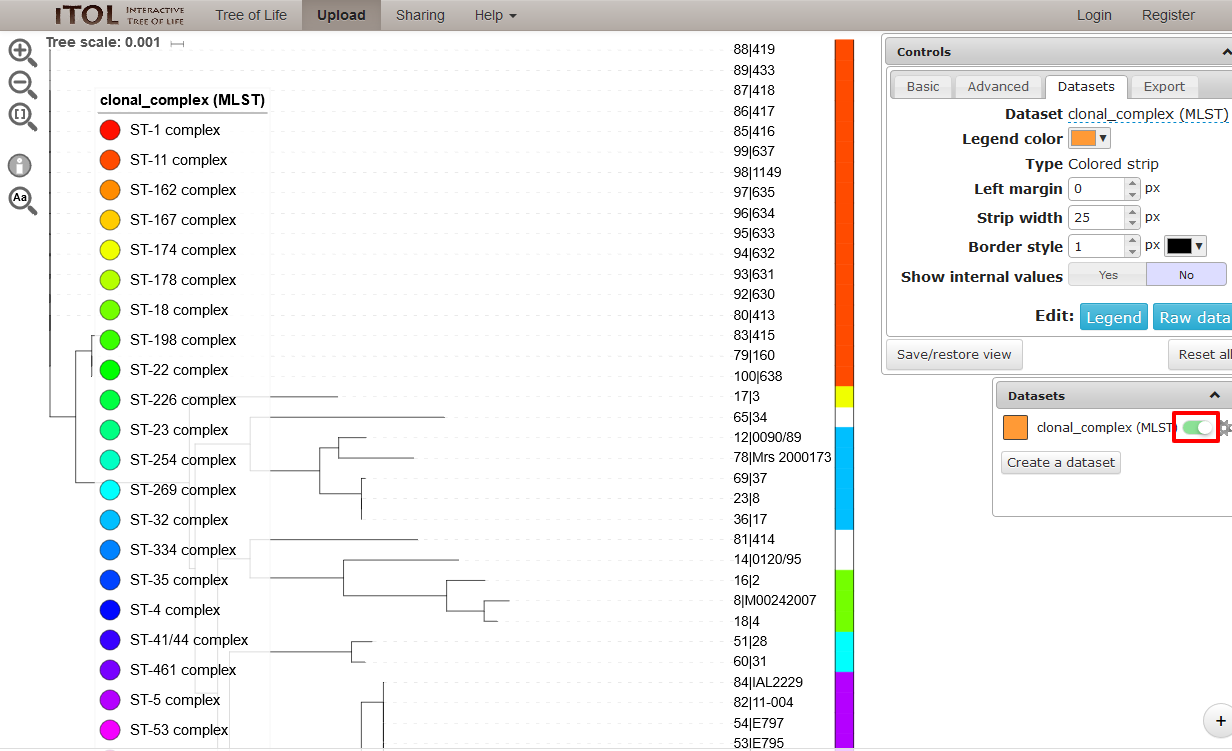
You can manipulate the tree in the browser, and display metadata by selecting the appropriate toggle.
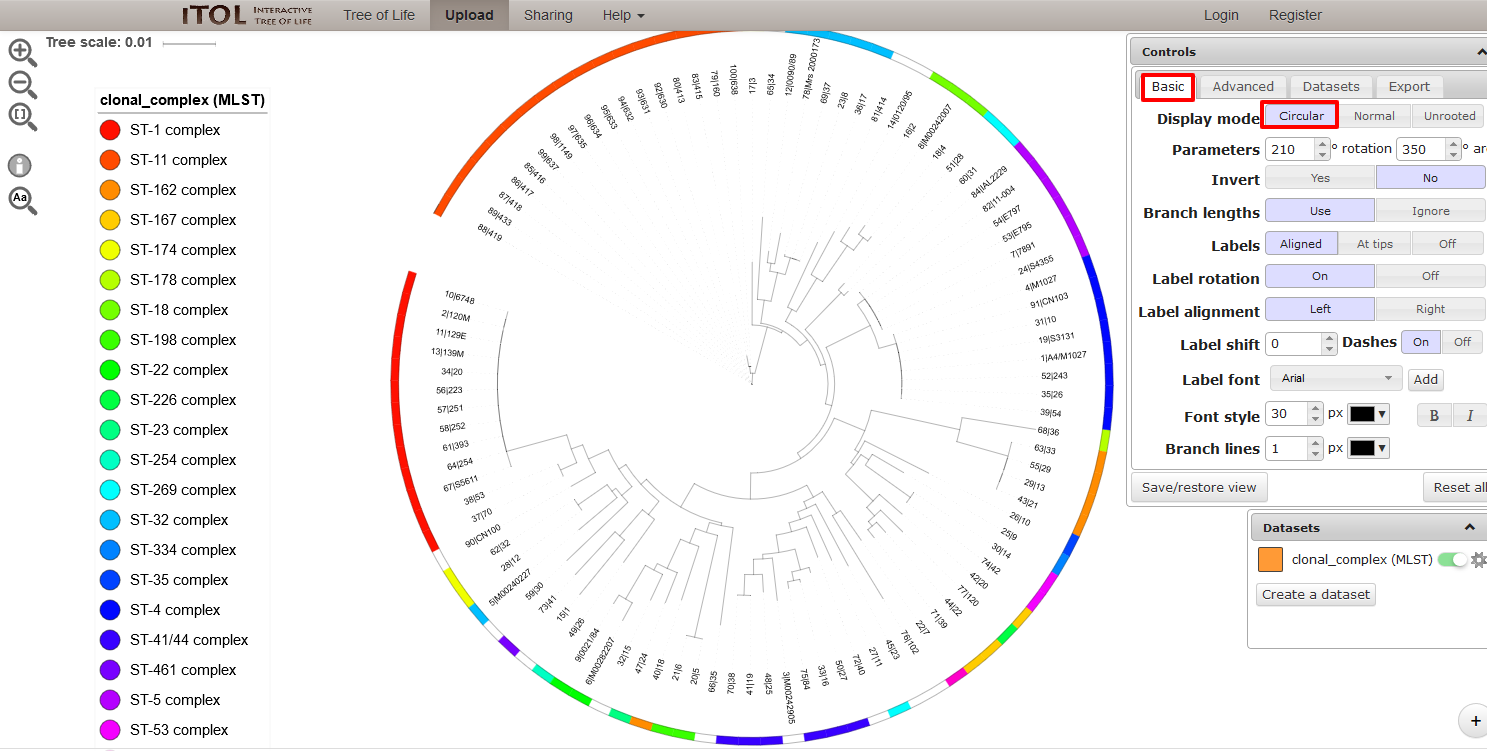
The tree layout can be changed by clicking the ‘Basic tab’ and, for example, selecting a circular display mode.
See the detailed documentation on the iTOL website for more information about manipulating and exporting trees.