BURST
BURST is an algorithm used to group MLST-type data based on a count of the number of profiles that match each other at specified numbers of loci. The analysis is available for both sequence definition database and isolate database schemes that have primary key fields set. The algorithm has to be specifically enabled by an administrator. Analysis is limited to 1000 or fewer records.
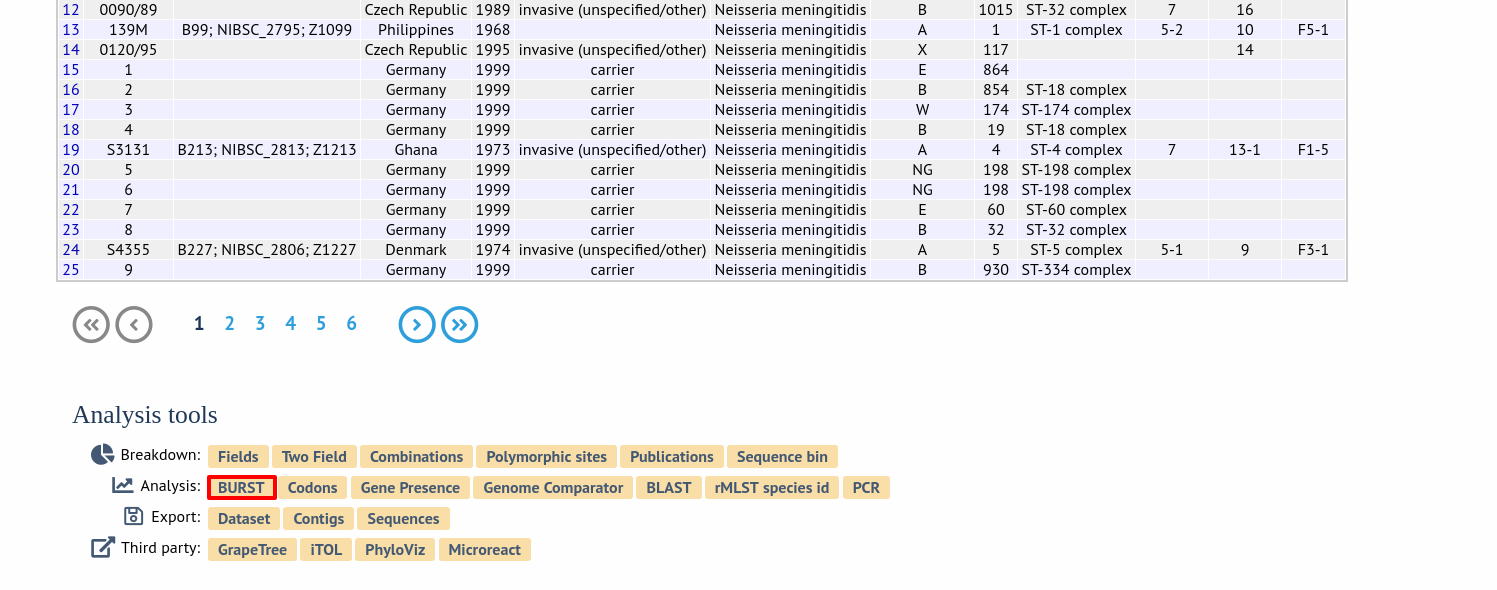
The plugin can be accessed following a query by clicking the ‘BURST’ button in the Analysis list at the bottom of the results table. Please note that the list of functions here may vary depending on the setup of the database.
If there multiple schemes that can be analysed, these can then be selected along with the group definition.
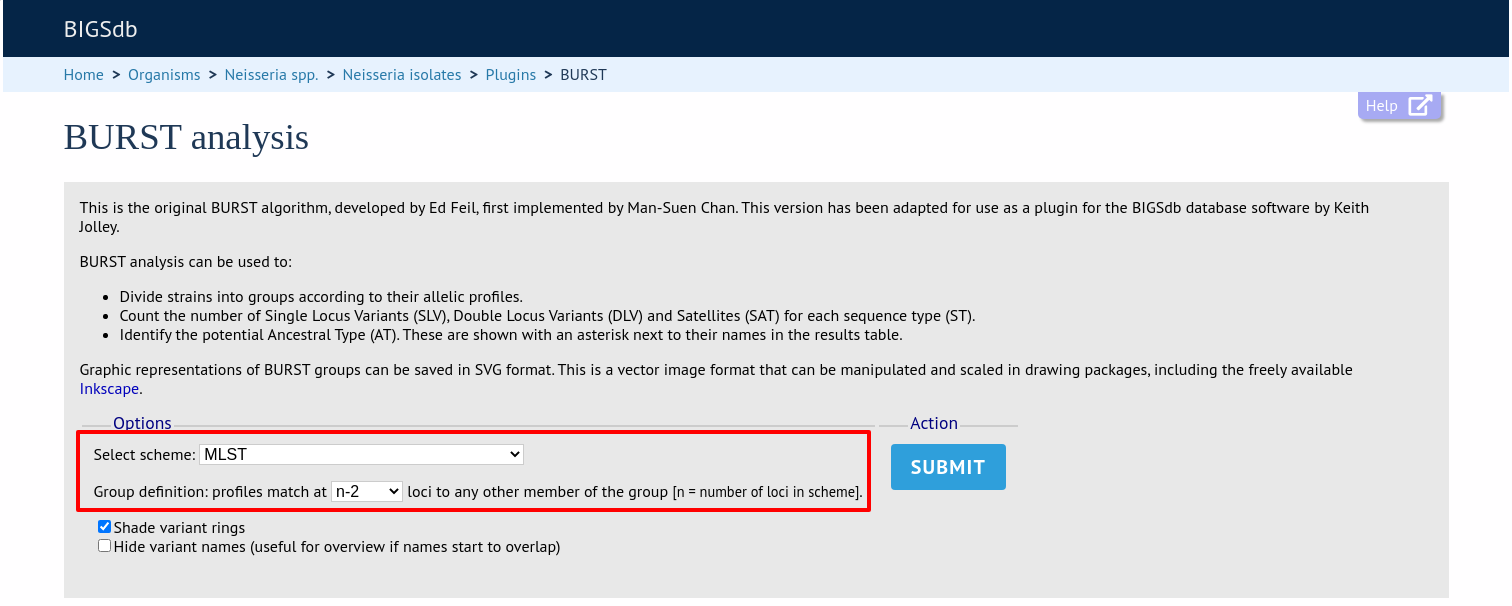
Modifying the group definition affects the size of groups and how they link together. By default, the definition is n-2 (where n is the number of loci), so for example on a 7 locus MLST scheme groups contain STs that match at 5 or more loci to any other member of the group.
Click Submit.
A series of tables will be displayed indicating the groups of profiles. Where one profile can be identified as a central genotype, i.e. the profile that has the greatest number of other profiles that are single locus variants (SLV), double locus variants (DLV) and so on, a graphical representation will be displayed. The central profile is indicated with an asterisk.
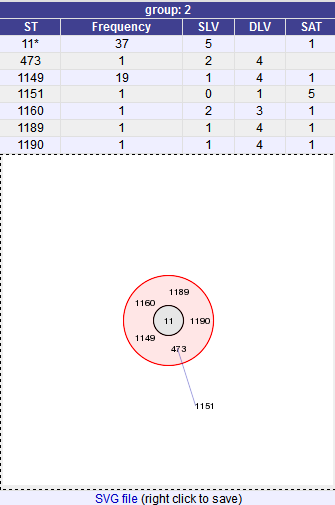
SLV profiles that match the central profile are shown within a red circle surrounding the central profile. Most distant profiles (triple locus variants) may be linked with a line. Larger groups may additionally have DLV profiles. These are shown in a blue circle.
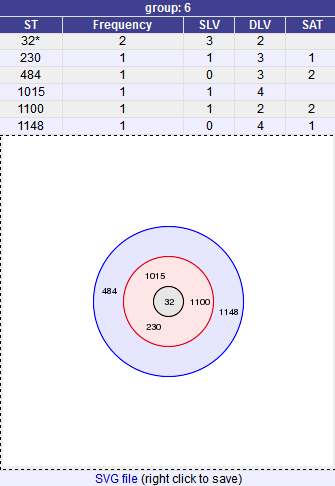
Groups can get very large, where linked profiles form sub-groups and an attempt is made to depict these.