Updating locus descriptions
Loci in the sequence definitions database can have a description associated with them. This may contain information about the gene product, the biochemical reaction it catalyzes, or publications providing more detailed information etc. This description is accessible from various pages within the interface such as an allele information page or from the allele download page.
Note
In recent versions of BIGSdb, a blank description record is created when a new locus is defined. The following instructions assume that this is the case. It is possible for this record to be deleted or it may never have existed if the locus was created using an old version of BIGSdb. If the record does not exist, it can be added by clicking the Add (+) button in the ‘locus descriptions’ box. Fill in the fields in the same way as described below.
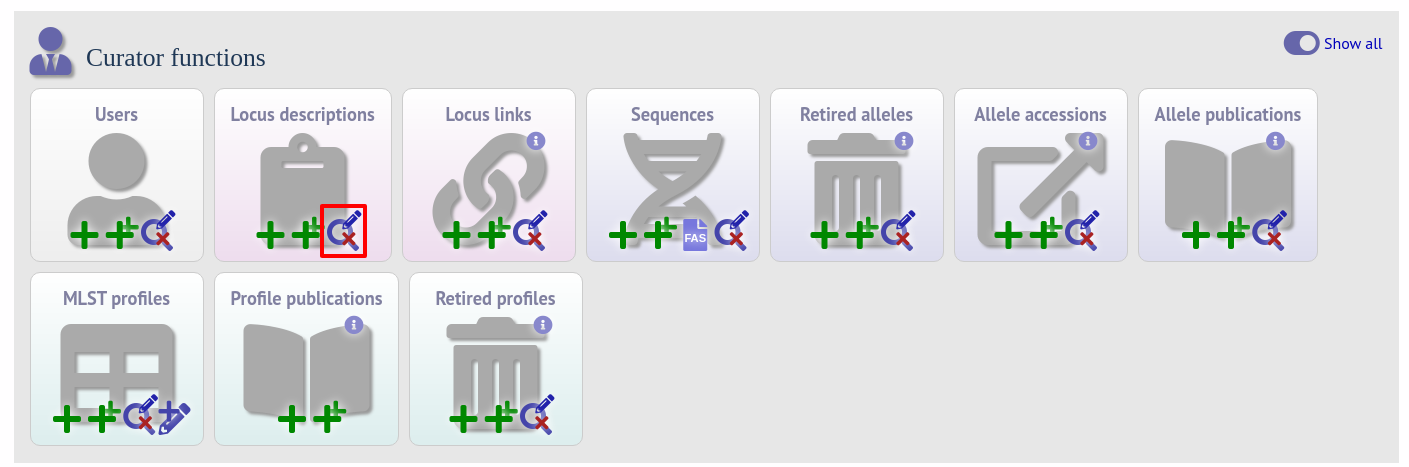
To edit a locus description, first you need to find it. Click the update/delete button in the ‘locus descriptions’ box on the sequence database curator’s page (depending on the permissions set for your user account not all the links shown here may be displayed). This function is normally hidden, so you may need to click the ‘Show all’ toggle to display it.
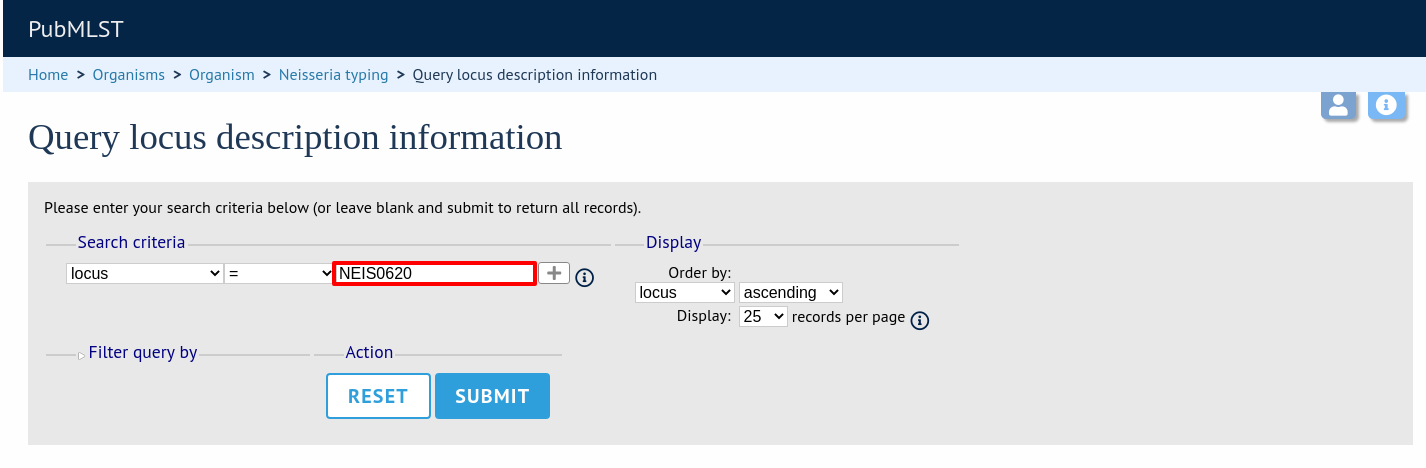
Either enter the name of the locus in the query box:
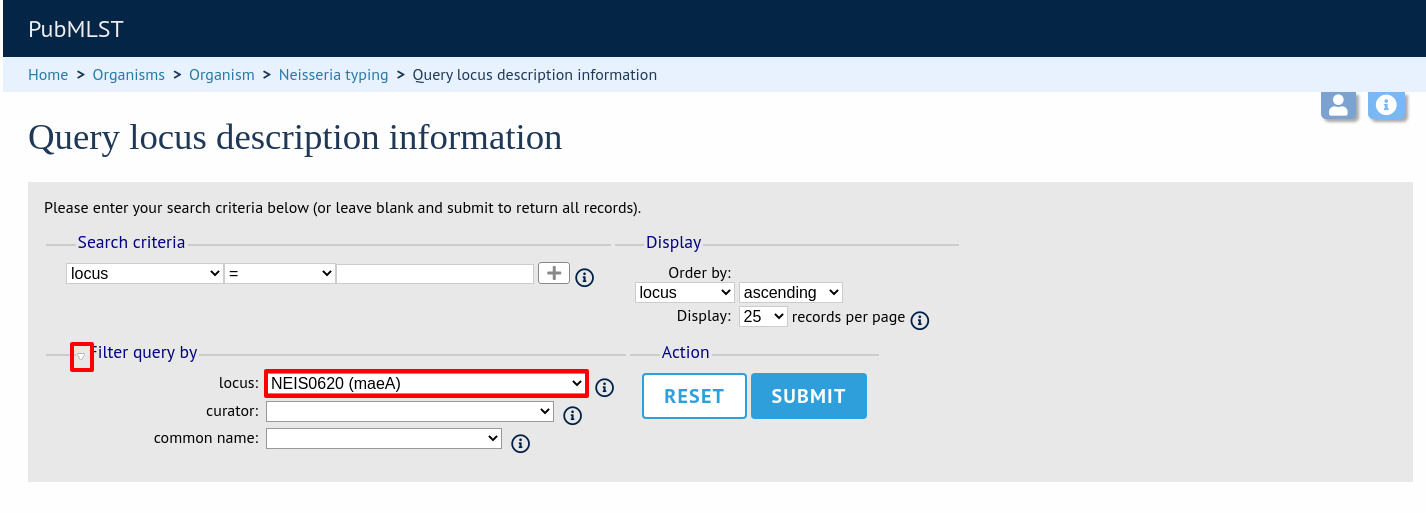
or expand the filter list and select it from the dropdown box:
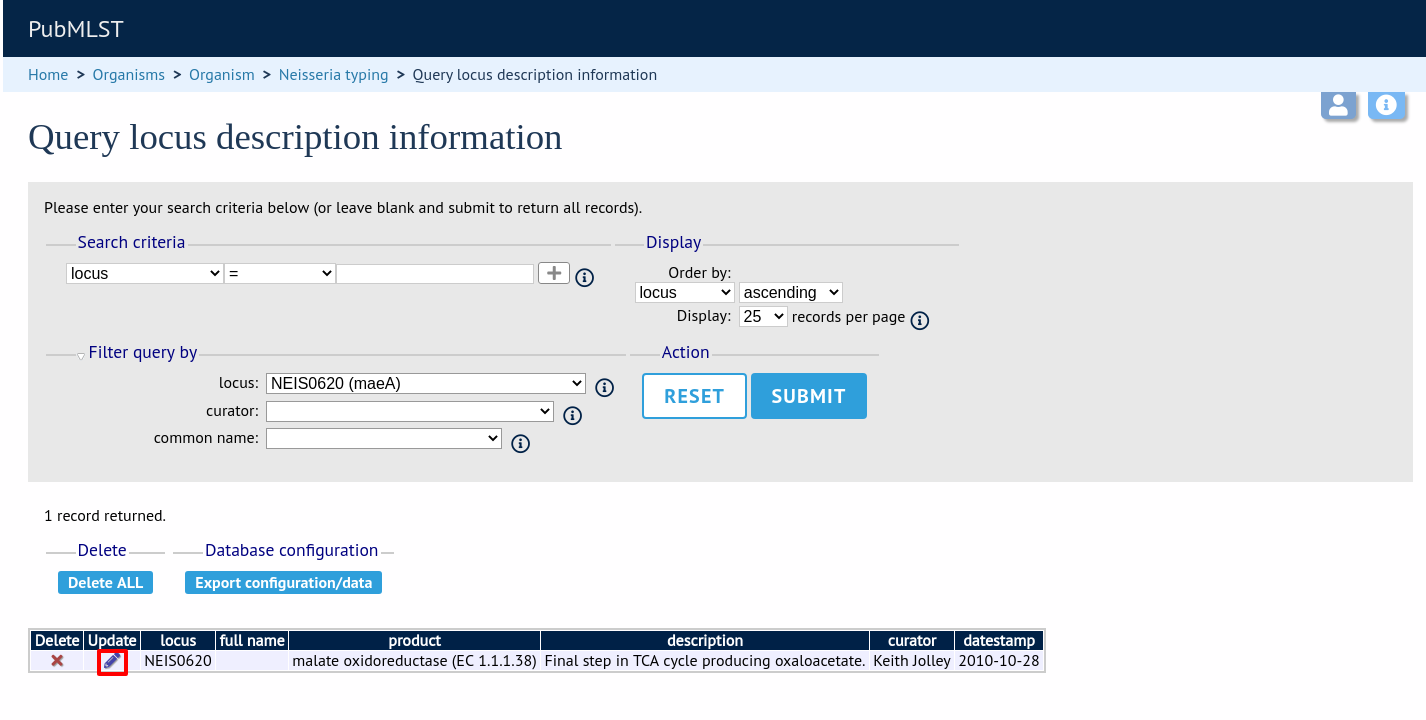
Click ‘Submit’.
If the locus description exists, click the ‘Update’ link (if it doesn’t, see the note above).
Fill in the form as needed:
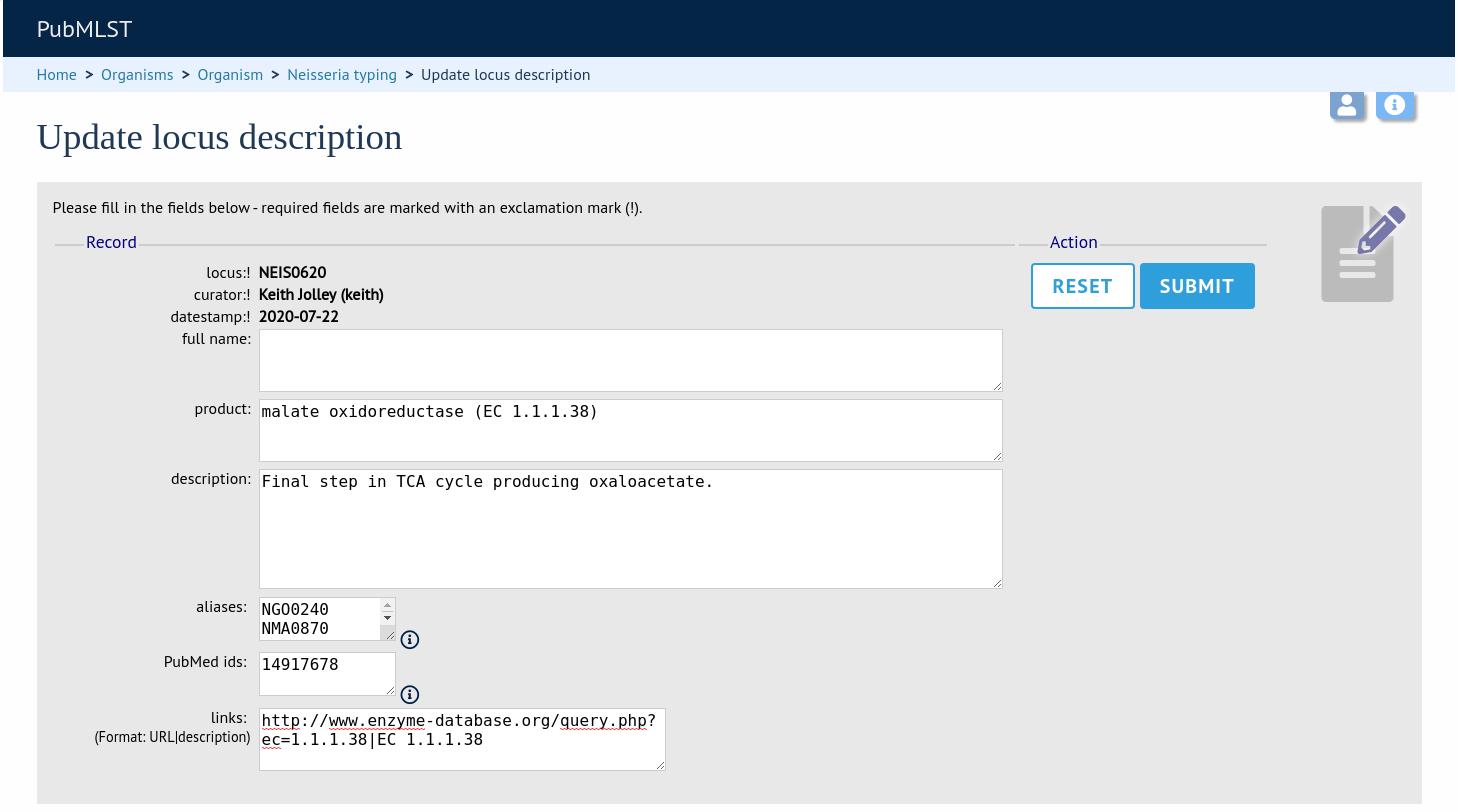
full_name
The full name of the locus - often this can be left blank as it may be the same as the locus name. An example of where it is appropriately used is where the locus name is an abbreviation, e.g. PorA_VR1 - here we could enter ‘PorA variable region 1’. This should not be used for the ‘common name’ of the locus (which is defined within the locus record itself) or the gene product.
product
The name of the protein product of a coding sequence locus.
description
This can be as full a description as possible. It can include the specific part of the biochemical pathway the gene product catalyses or may provide background information, as appropriate.
aliases
These are alternative names for the locus as perhaps found in different genome annotations. Don’t duplicate the locus name or common name defined in the locus record. Enter each alias on a separate line.
Pubmed_ids
Enter the PubMed id of any paper that specifically describes the locus. Enter each id on a separate line. The software will retrieve the full citation from PubMed (this happens periodically so it may not be available for display immediately).
Links
Enter links to additional web-based resources. Enter the URL first followed by a pipe symbol (|) and then the description.
Click ‘Submit’ when finished.